RICHLAND, Wash., May 2, 2006 – Scientists at Pacific Northwest National Laboratory have used electrostatic attraction to layer reactive biological molecules in a lasagna-like way around spaghetti-like carbon nanotubes. The configuration can be used in a wide range of applications, from ultraprecise blood sugar monitoring to infectious agent detection, said Yuehe Lin, who led the research at the Department of Energy campus’ W.R. Wiley Environmental Molecular Sciences Laboratory.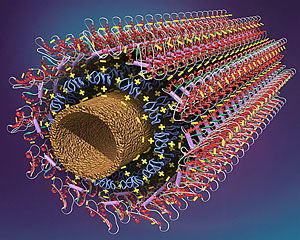
A carbon nanotube. (Images: William R. Wiley Environmental Molecular Sciences Laboratory)
The technique, described in the April Journal of Nanoscience and Nanotechnology, enables enzymes, with the help of a long, noodle-like polymer molecule, to self-assemble layer-by-layer on a single carbon nanotube. Lin and co-author Guodong Liu, a postdoctoral fellow in Lin’s group, coaxed electrostatic clinginess in a polymer and an oppositely charged protein-enzyme, in this case glucose oxidase, which reacts in the presence of blood sugar. The catalyzed products from the reaction "ping" the carbon nanotube; if the tube is connected to an electrode, the tube will carry a signal that corresponds precisely with the amount of glucose detected. The first polymer binds to the carbon nanotube. Enzymes are attracted to the polymer, leaving an outer layer for the next polymer of opposite charge to cling to, and so on. 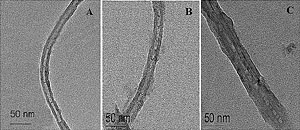
Electron-microscopic images show enzyme-coated carbon nanotubes of varying thicknesses. The sensitivity of a biosensor increases with thickness, to about six layers (far right).
An individual strand coated with one to six layers ranges from 30 nm to about 50 nm thick. “Each polymer layers is porous,” Lin said. “This allows the glucose to diffuse in and come into contact with the enzymes.”
The polymers trap the enzymes in place,” Liu said. “You can go up to five or six layers to improve the sensitivity of the detector, but after that, the more enzyme layers” inhibit diffusion of the glucose.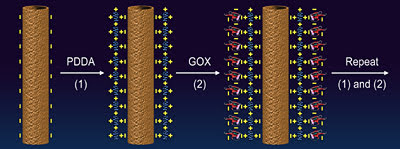
A polymer, here labeled PDDA (far left), clings to a carbon nanotube of opposite charge, and an enzyme, GOX, does the same with the polymer. The steps can be repeated to build up a biosensor's layers, enzyme count and sensitivity.
Now that the glucose enzyme biosensor has passed the test, Lin said, it should be possible to build a similar sensor using other enzymes that react specifically with other biological chemicals, environmental pollutants or even microbes and their toxic byproducts. Lin and colleagues also reported, in February issue of Analytical Chemistry, having used a similar technique to construct sensors for nerve agents.
The research was funded through PNNL’s environmental biomarker initiative. For more information, visit: www.pnl.gov