Despite the critical role our brains play in perception, cognition and action, we know very little about how the brain actually accomplishes these feats. With the advent of new optical methods for probing neural networks, researchers have begun to untangle the neural web and shed new light on the mysteries of the brain. These optical techniques are now being taken a step further with the incorporation of spatial light modulators, or SLMs, that enable parallel recording and control of larger neural populations by generating dynamic 3D holograms inside of living brains.
DR. CHRISTOPHER HOY, BOULDER NONLINEAR SYSTEMS INC.
The brain truly is a marvelous wonder of biology. With an extensive range of functions and dynamics that span many length scales and timescales, it is also marvelously complex. By better understanding our brains’ neural networks, researchers hope to improve our understanding of neurological diseases, such as Alzheimer’s and schizophrenia. This understanding also will advance our knowledge of computing technologies and brain-machine interfaces and help elucidate seemingly intractable mysteries including memory, dreaming and consciousness.
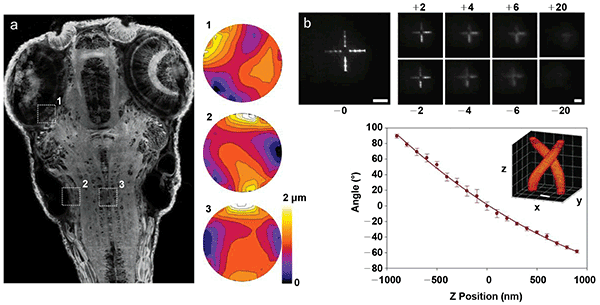
Additional microscope functionality provided by spatial light modulators (SLMs). (a) Adaptive aberration correction in two-photon fluorescence maximum intensity projection of living zebrafish brain. The numbered wavefront corrections to the right of the image correspond to numbered regions in image and demonstrate the variability of the aberration correction required in different parts of the image. Courtesy of K. Wang and E. Betzig/Janelia Research Campus. (b) Nonmechanical remote focusing of a test pattern generated by two-photon-excited fluorescence away from the focal plane of the imaging camera. Scale bars are 50 μm(3). Courtesy of R. Yuste/Columbia University. (c) Superresolution localization through the use of an engineered, double-helix point spread function. Scale bar is 400 nm7. Courtesy of W.E. Moerner/Stanford University.
The critical structures and processes in the brain range from molecular-scale neurotransmitter interactions up through cellular and multicellular structures and responses. In turn, these interactions coordinate the activity of the whole brain as an organ, ultimately resulting in the behavior of the animal or human. Similarly, critical timescales in the brain range from the millisecond scale of single action potentials all the way up to the multiday timescales associated with learning and memory. As a result, the field of neuroscience is inherently “multiscale,” and multiscale phenomena are notoriously difficult to understand and model.
Recently, research in this field has intensified due to numerous high-profile initiatives such as the U.S. National Institutes of Health’s (NIH) Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Initiative (related to both the Brain Activity Map (BAM) Project and the Human Connectome Project), the European Commission’s Human Brain Project, and Japan’s Brain Mapping by Integrated Neurotechnologies for Disease Studies (Brain/MINDS) Project. Notably, BRAIN, BAM and Brain/MINDS have each identified the critical need for improving the optical technologies used for the functional mapping of neural networks.
Conventional whole-brain imaging techniques, such as functional magnetic resonance imaging (fMRI) and diffusion tensor imaging (DTI), are powerful tools for visualizing neural activity and connections across regions of the brain. Their spatial resolutions, however, are limited to the millimeter scale and therefore they cannot resolve the individual neurons that make up the fundamental functional units of the brain. The fMRI and DTI methodologies also do not image action potentials or neural connections directly, but rather rely upon surrogate signals (i.e., blood flow and water diffusion, respectively). At the other spatiotemporal extreme, single-cell electrophysiology measurements can directly record the action potentials of individual neurons via microelectrodes. However, this approach is inherently invasive and limited in the number of neurons that ultimately can be observed in a given experiment. It is here where optics come into play.
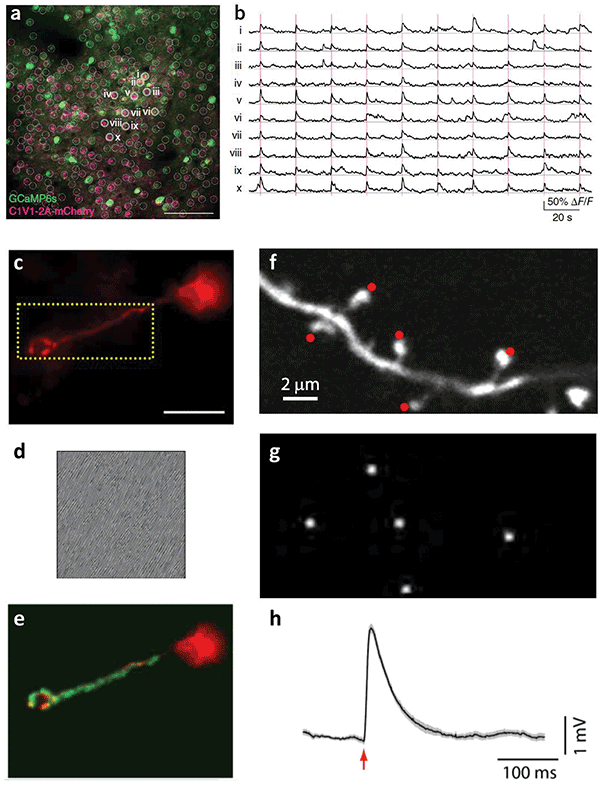
Holographic targeting of neurons. (a) Simultaneous photostimulation of neuron subpopulation and (b) Concurrent optical recording of induce calcium transients4. Courtesy of M. Haüsser/University College London. (c) Targeting of a single dendrite with (d) the corresponding phase mask displayed on the spatial light modulator and (e) the subsequently shaped illumination in green2. Courtesy of V. Emiliani/Neurophotonics Laboratory, CNRS, Paris Descartes University. (f) Targeting of multiple dendritic spines for glutamate uncaging with (g) patterned illumination and (h) the average uncaging potential as measured by whole-cell patch clamp recording3. Courtesy of R. Yuste/Columbia University.
Optical technologies compliment these existing tools by providing direct, noninvasive imaging of neurons and their respective action potentials with single-cell resolution across small submillimeter volumes. Furthermore, the use of photostimulation and photoinhibition gives neuroscientists the ability to either induce or inhibit action potentials in neurons, going beyond simply listening to neural activity to actually directing it. This ability to perturb local neural networks while simultaneously observing the result is critical to improving our understanding of neural circuitry.
On the biology side, advances in optical techniques for functional connectomics have been led by the development of extrinsic voltage- or calcium-sensitive fluorophores and photoactivation compounds, as well as by the development of their intrinsic analogs through optogenetics. Voltage- and calcium-sensitive fluorophores modulate their respective fluorescence with changes in membrane voltage or calcium ion concentration to allow visualization of individual action potentials. Optogenetics takes this a step further by genetically encoding targeted neurons to express similar fluorescent proteins and light-sensitive ion channels for direct reporting and control of action potentials. Through optogenetics, neuro-scientists are developing a complete toolbox for visualizing networks, manipulating neural activity, and recording the result—an ideal suite of capabilities for functional brain mapping over small length scales ranging on the orders of 0.1 to 1.0 mm3.
If the promise of optogenetics is to noninvasively bridge the gap between single-cell electrophysiology and whole-brain imaging techniques, then new optical technologies are needed to enable simultaneous stimulation and the recording of larger populations of neurons within intact neural circuits. In fact, it is the correlated activity of neuron groups, termed “ensembles,” that are believed to be significant for a wide range of brain functions, ranging from motor activity and navigation to learning and memory.1
Groups of genetically identical neurons can be stimulated and recorded optogenetically en masse using diffuse illumination (e.g., via a fiber optic probe). However, the study of subpopulations within populations of genetically identical neurons requires precise cellular targeting. Furthermore, the targeted stimulation and recording of neurons at depths below the most superficial hundreds of microns requires the use of deeply penetrating, near-infrared (NIR) light and the simultaneous absorption of two or three photons using multiphoton microscopy. Because the nonlinear nature of multiphoton microscopy requires focused light, the simultaneous investigation of neuron groups requires the generation of many independent, focused “beamlets.” In 2008, Lutz et al. and Nikolenko et al. independently demonstrated that this can be achieved using a phase-modulating spatial light modulator (SLM) to create 3D-patterned light through digital holography in both single-photon and multiphoton regimes, respectively.2, 3
The SLMs commonly used in digital holography are pixelated, liquid crystal on silicon (LCOS) devices akin to the LCDs used for computers and televisions. In a phase-modulating spatial light modulator, termed an LCOS-SLM, the pixel voltage is used to rotate birefringent nematic liquid crystal molecules, resulting in a change in the local refractive index and thus a programmable phase delay at each pixel. In digital holography, the SLM is used to impose a user-defined wavefront on the illumination light with the SLM imaged to the back aperture of the microscope’s objective lens. The pattern on the SLM can be simple, such as a blazed grating to laterally steer the focus at the sample or a Fresnel lens to axially steer the focus. The pattern can also be incredibly complex, providing holographic generation of 3D arrays of focal spots or even arbitrary shapes and patterns.
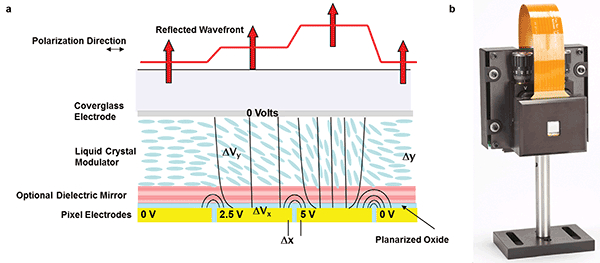
Phase modulation with a liquid crystal on silicon spatial light modulator (LCOS-SLM). (a) Schematic of spatial light modulator showing the response of the liquid crystal to electric field and subsequent shaping of the wavefront. (b) Photo of a commercial liquid crystal spatial light modulator. Courtesy of Meadowlark Optics.
The speed of LCOS-SLMs is limited largely by the transient response time of the liquid crystal molecules to the applied voltage and by the rate at which pixel voltages can be updated. While many SLMs have emerged from the consumer display industry and operate at 60 Hz, the fastest commercial SLMs can now modulate the NIR light used in multiphoton optogenetics at frame rates exceeding 200 Hz. Originally developed by Boulder Nonlinear Systems (BNS) and now produced and sold by Meadowlark Optics, these devices enable dynamic holographic patterning in multiphoton microscopes at speeds approaching the single-millisecond timescale of individual action potentials.
Using current LCOS-SLMs, researchers are able to simultaneously stimulate and record functional ensembles of neurons4 or parts of a single neuron, such as an elongated dendrite2 or individual dendritic spines.3,5 These types of experiments are critical to understanding neural dynamics at the ensemble- and single-cellular levels, respectively, and cannot be achieved with optogenetic labeling alone. Meanwhile, because the SLMs can quickly and nonmechanically simulate arbitrary optical transfer functions, SLM-equipped microscopes provide additional functionality beyond holographic cellular targeting and light shaping. In fact, SLMs represent a sort of optical “Swiss Army Knife” for laser scanning microscopes, capable of providing adaptive optics for deeper imaging,3 remote focusing for nonmechanical 3D imaging,3 or engineered point-spread functions for superresolution localization.7 According to holographic neuroscience pioneer and Columbia University professor Rafael Yuste, who serves as Co-Director of Columbia’s Kavli Institute for Brain Science, “SLMs give us the ultimate flexibility to craft the light in space and time.”
Despite the many advantages SLMs bring to laser scanning microscopes, their adoption by biologists remains limited in part by the technical hurdles of hardware and software integration. This integration frequently requires modifications to expensive commercial microscopes and custom software development that often provides limited, if any, communication with the native microscope software. To tackle this challenge, BNS teamed up with the Yuste Lab in 2012 to conduct a two-phase development project funded by the National Institute for Mental Health (NIMH) to bring turn-key SLM microscopy solutions to the neuroscience market.
In the first phase, BNS and the Yuste Lab commercially developed a portable single-photon SLM microscope that previously had been developed by the lab.8 This small but versatile platform, dubbed the “Pocketscope,” is capable of in vitro, holographic multicell recording of action potentials within brain slices.9 The Pocketscope measures 22 × 42.5 × 30 cm and includes a 512 × 512-pixel LCOS-SLM, as well as a continuous wave (CW) 488-nm laser and white LED for either holographically patterned epifluorescence or conventional brightfield imaging. Most importantly, the Pocketscope is driven by a stand-alone user interface that fully integrates the SLM control and hologram calculation into the imaging functionality of the microscope — this provides seamless holographic light patterning and aberration correction. With this software, real-time holographic patterns can be generated automatically through either an automated cell body detection feature or an intuitive point-and-click targeting interface. The software also offers the ability to record images while cycling through sequences of holographic patterns. In this manner, the Pocketscope is able to target tens of cells simultaneously throughout a 440 × 440-μm field of view. Currently, the system is in premarket refinement prior to commercial release.
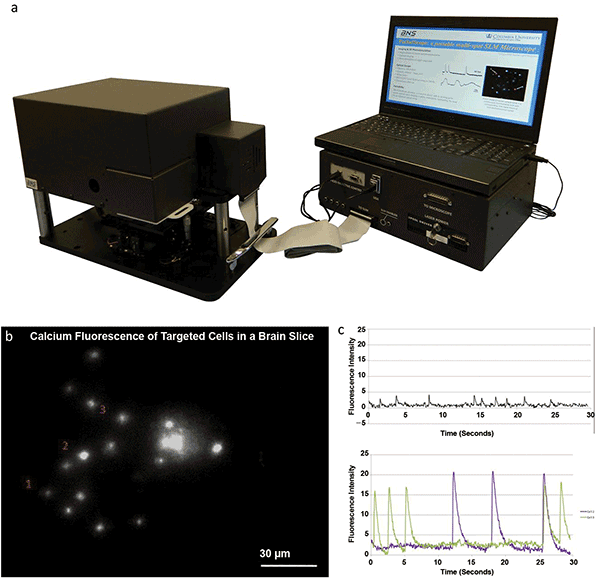
The Pocketscope, a turn-key SLM microscope for neuroscience applications. (a) Photo of the Pocketscope system commercially developed by Boulder Nonlinear Systems (BNS) and Columbia University’s Yuste Lab. Courtesy of BNS. (b) Frame from a 30-second recording of holographic calcium imaging of 15 neurons in a brain slice with (c) recorded calcium transients from labeled cells 1, 2 and 3 (black, purple and green traces, respectively)9. Courtesy of A. Linnenberger.
The Pocketscope’s use of conventional single-photon excitation is well-suited for the investigation of thin in vitro samples. However, many phenomena only can be investigated through the study of neural circuitry deep in the brains of awake, behaving animals. In order to accomplish this, the deep penetration and 3D precision of two- or three-photon excitation are required. Currently in the second phase, BNS and the Yuste Lab are addressing this need by developing a bolt-on SLM module for commercial multiphoton microscopes in partnership with Meadowlark Optics.
The development of SLM-based systems for the neuroscience community parallels ongoing developments to improve the SLMs themselves. The ideal experimental parameter regimes for holographically patterned optogenetics are not yet well-known, due largely to existing hardware limitations. The ideal SLM would be capable of generating diffraction-limited focal spots anywhere in the microscope’s field of view. Furthermore, the SLM would also be capable of switching at speeds exceeding the millisecond timescale of neural circuits so as to enable closed-loop experiments. To enable researchers to use the best wavelength for a given optogenetic target, the SLM must also modulate efficiently across a broad wavelength range compatible with tunable
laser sources. Finally, the ideal SLM would be able to handle the highest laser powers for efficient multiphoton excitation across hundreds of independent spots. That’s a tall order for existing devices. However, current research suggests that these capabilities soon may be available.
For example, SLMs employing ferroelectric liquid crystal (as opposed to the nematic liquid crystal, which is commonly employed in phase-modulating SLMs) can provide 1-kHz frame rates and broadband modulation when used as a phase modulator. Such devices, though, are limited in their modulation ranges to less than a full wave of phase, and existing commercial devices provide only binary modulation. By using analog modulation of a high-tilt ferroelectric liquid crystal, we hope to address both of these issues through the National Institutes of Health- (NIH) funded effort to develop a high-speed achromatic SLM. This component-level development effort complements another recent NIH-awarded investigation that takes a holistic approach to designing the next generation of optogenetic SLMs. This investigation, which is being undertaken in collaboration with MIT’s professor Ed Boyden, who helms the institute’s Synthetic Neurobiology Group, aims to develop new SLM modeling capabilities to guide the development of SLMs based on the simulated performance of new devices in existing multiphoton microscopes.
The continued developments of SLM components and systems aim to not only expand the capabilities of holographically patterned optogenetics, but to also expand the adoption of these capabilities in the wider neuroscience community. With the added flexibility SLMs provide to optical microscopes, neuroscientists hopefully will be one step closer to a day when they are limited not by their hardware but only by their imaginations.
Meet the author
Dr. Christopher Hoy is Research Scientist at Boulder Nonlinear Systems in Lafayette, Colo.; email: [email protected].
References
1. B. Haider and D.A. McCormick (2009). Rapid neocortical dynamics: cellular and network mechanisms. Neuron, Vol. 62, pp. 171-189.
2. C. Lutz, T.S. Otis et al. (2008). Holographic photolysis of caged neurotransmitters. Nat Methods, Vol. 5, pp. 821-827.
3. V. Nikolenko, B.O. Watson et al. (2008). SLM microscopy: scanless two-photon imaging and photostimulation with spatial light modulators. Front Neural Circuits, Vol. 2.
4. A. Packer, L. Russell et al. (2015). Simultaneous all-optical manipulation and recording of neural circuit activity with cellular resolution in vivo. Nat Methods, Vol. 12, pp. 140-146.
5. S. Yang, E. Papagiakoumou et al. (2011). Three-dimensional holographic photostimulation of the dendritic arbor. J Neural Eng, Vol. 9, p. 046002.
6. K. Wang, D.E. Milkie et al. (2014). Rapid adaptive optical recovery of optimal resolution over large volumes. Nat Methods, Vol. 11, pp. 625-628.
7. S. Pavani, M. Thompson et al. (2009). Three-dimensional, single-molecule fluorescence imaging beyond the diffraction limit by using a double-helix point spread function. Proc Natl Acad Sci, Vol. 106, pp. 2995-2999.
8. V. Nikolenko, D.S. Peterka et al. (2010). A portable laser photostimulation and imaging microscope. J Neural Eng, Vol. 7, p. 045001.
9. A. Linnenberger, D.S. Peterka et al. (2014). The Pocketscope: a spatial light modulator based epi-fluorescence microscope for optogenetics. Proceedings of SPIE NanoScience + Engineering, p. 91640Y-91640Y.