In recent years, optical tweezers have generated remarkable interest in research areas outside of fundamental physics, including biochemistry, biology and medicine. Modern tweezer designs enable researchers from all disciplines to benefit from the advantages of this technique.
VITALIY OLIYNYK, PHILIPP RAUCH AND STEFAN B. KAEMMER, JPK INSTRUMENTS
Optical tweezers offer a noninvasive experimental method to trap, track and manipulate microscopic objects ranging from several tens of micrometers to tens of nanometers in size. The trapped particles can be accurately measured and manipulated with high temporal, spatial and force resolution. Experiments can show spatial resolution in the nanometer range while obtaining force data with piconewton resolution on a millisecond timescale. The basic physical phenomenon underlying optical tweezers is the momentum transfer of light, when it collides with matter. Due to the noninvasive nature and the capability to work under physiologically relevant conditions, optical tweezers have found widespread applications in cell biology and medical research to study chemical and physical processes, from biological polymers and molecular machines to the manipulation of single viruses and bacteria.
Recent discoveries
Since the pioneering work of Arthur Ashkin1 30 years ago, applications for optical tweezers have grown tremendously and have led to a number of scientific discoveries. Optical tweezer approaches have been extended down to the single-molecule level and have made fundamental contributions to the detailed understanding of the mechanochemistry of molecular motors, which power muscles and facilitate transport of material on a cellular level. As an example, optical trapping was used to measure the 8-nm steps taken by kinesin, a motor protein. The steps were quantified as the molecule walked along a microtubule and it was shown that such motors advance progressively over very long distances encompassing hundreds of single steps2 (Figure 1). Other remarkable discoveries aided by the use of optical tweezers include investigations of the bacterial version of RNA polymerase3; the motions of ribosome, a giant macromolecular machine composed of proteins and nucleic acids4; the motor proteins myosin5 and dynein6; and the directed force generation by cross-linkers in microtubule networks7.
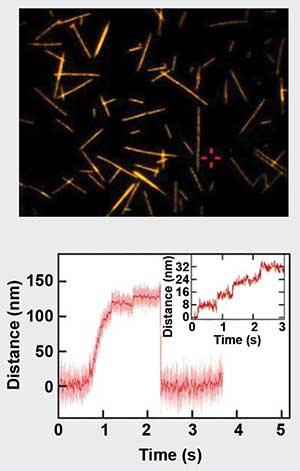
Figure 1. Motility assay of stepping motor-protein Kinesin-1. On the top is a fluorescence microscopy image of microtubules on glass coverslips. The lower panel depicts the force generated by a kinesin molecule, which walks along the microtubules in 8-nm steps. Sample courtesy of Dr. Erik Schaeffer/TU Dresden.
Another important area of research includes the study of biopolymers, such as DNA and its interactions with associated proteins. These proteins affect the structure of DNA, and thus the force-dependent length of the DNA molecules (Figure 2). In optical tweezers, these length changes, as well as the kinetics of the protein association and dissociation, can be observed with a high degree of accuracy8,9.
The last decade has seen much of the momentum for applications of optical tweezers in single-cell and medical research. The reason for this trend is twofold: 1. the ability of optical tweezers to measure forces and distances in the piconewton and nanometer range, which matches the requirements for studying cellular interactions at the single molecule level; and 2. that optical trapping, even inside a living cell, can be performed almost noninvasively. During the past decade, tweezers were successfully used to reveal binding and unbinding pathways of a single influenza virus to its host cell10 and to study force guided cell motility11. Tweezers were also used to look at stem cell differentiation12, reveal the mechanisms of fast recycling of plasma membranes and vesicular proteins required for the sustained exocytosis13;, and to push the development of nanomaterial-mediated therapies for treatment of tumors14.
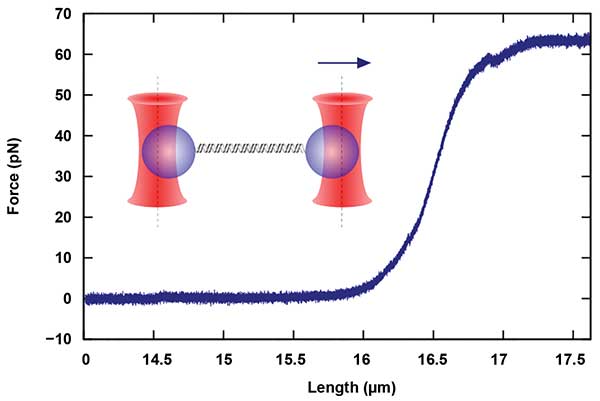
Figure 2. Measurements of single-molecule mechanics, in this case, a double-stranded DNA (dsDNA) molecule, are performed with dual-beam devices that allow the parallel manipulation and force detection with two independent traps. The molecule termini are chemically attached to two different particles. By moving one of the traps, a force is applied to the molecule. Courtesy of JPK Instruments AG.
Technical advances
Easily accessible and powerful, modern optical tweezers are built on top of commercial high-end inverted optical microscopes utilizing immersion objectives with high numerical apertures (NA), precise sample translation stages, and fast, accurate beam steering methods (Figure 3). The minimum viable configuration consists of a microscope with a coupled laser beam that is focused into a sample chamber with the help of a high-NA objective. During the two decades of development the instruments went a long way from such minimalistic configuration of having a single static trap to configurations with multiple independently steerable traps, reliable force detection and particle tracking schemes. A general approach to building the most universal tweezer setup covering the greatest number of biochemical and biophysical applications has been realized. Today, optical tweezers vary in the method to steer and/or split the laser beam, the design of the force measurement and particle tracking scheme, as well as the data acquisition electronics and software driving the various experiments and the accessories to enable, for example, experiments on living matter.
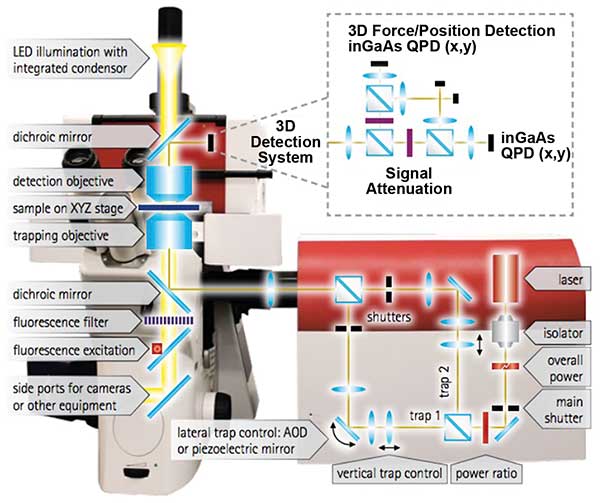
Figure 3. JPK Instrument’s NanoTracker 2 is an integrated optical tweezers system. The optics unit in the center contains the laser and optical elements for beam deflection and alignment. Courtesy of JPK Instruments.
Trap generation
JPK Instrument AG’s NanoTracker 2 implements the most widespread scheme in the optical tweezer field (Figure 3). This scheme involves a dual-trap setup based on an interferometric method for force measurement using an infrared solid-state laser source. The introduction of two traps, which are a requirement for a vast amount of bio-applications, such as studies of DNA mechanics or protein folding/unfolding experiments, can be done by using a single laser source and splitting the beam based on polarization into two. This approach reduces costs due to the lack of a second laser system while still allowing researchers to steer both beams independently away from or toward each other. Currently, at least two types of the beam steering systems can be used as well as their combination. The decision as to which system to use will be dictated by the experiment’s requirements.
For the first approach, a mirror is placed in the Fourier plane of the sample. To do this, the beam is tilted and the pivot point is imaged onto the back aperture of the microscope objective. The mirror is tilted normally in two orthogonal directions with the help of piezoelectric actuators resulting in a 2D motion of the trap inside the sample chamber. Such piezo-mirrors function with a closed-loop feedback providing nanometer precision of the trap positioning and relocation speeds close to 1 kHz.
The second approach is used in situations where very rapid movements of the traps are required. An entirely nonmechanical beam steering can be realized by acousto-optical deflectors (AODs). AODs utilize an acoustic wave propagating through a crystal and forming a diffraction grating due to pressure compression and depressions. When driven with suitable radio frequency (RF) synthesis electronics, this allows the researcher to modify the diffraction of the laser beam passing through the crystal. As a result, the speed of the beam steering is only limited by the sound velocity in the crystal, which leads to the ability of the AODs to relocate the laser beam at rates above 50 kHz. This feature allows the researcher to generate a significant number of traps with a single beam. Basically, the number of traps is limited only by the laser power. A beam driven by AODs can be scanned rapidly between several trap sites, such that particles do not have time to diffuse away between visits by the laser. In addition to simply manipulating multiple traps, the application of such trap multiplexing in conjunction with a suitably fast detection method allows researchers to measure the forces each trap “feels” by careful time synchronization with the scanner movement (Figure 4).
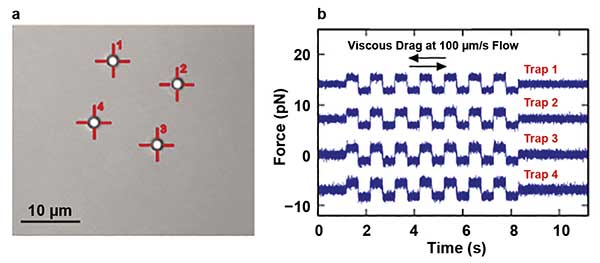
Figure 4. A bright-field image of four polystyrene 2-µm beads held by four traps is shown. The traps were created from the single laser source by the laser time-sharing principle using AOD scanners (a). Also shown are force measurements obtained from four multiplexed traps during a viscous drag experiment, where a piezo was oscillating with a constant speed of 100 µm/s (b). Courtesy of JPK Instruments.
Trapping laser
Diode-pumped, solid-state Nd:YAG (neodymium-doped yttrium aluminum garnet) lasers have become a typical choice for optical tweezers due to their cost-effectiveness, stability and high output powers. An additional advantage for investigating aqueous samples common in the biological field is that the Nd:YAG laser, with its 1064-nm wavelength, is close to the water’s absorption minimum. Consequently, the laser will only induce minimal sample heating, typically on the order of 1 ºC, and allow experiments with live cells to continue for hours at a time. Additionally, the use of an IR laser greatly facilitates the combination of optical tweezing with other microscopic methods in the same setup. Common optical imaging techniques, such as epi-fluorescence, total internal reflection fluorescence and confocal microscopy, can be carried out simultaneously as they operate predominantly in the visible spectrum of light.
Force detection
In the last decade, interferometric tracking of the trapped particles’ positions has evolved as the de facto standard for quantifying the trapping forces over other methods such as video-based tracking. The interferometric scheme involves a detector such as a quadrant photodiode (QPD) placed in the back focal plane of the microscope’s condenser lens, such that the direction of the light that has passed the trapped object determines the position of the beam on the QPD (Figure 3). A high-NA condenser is required to collect most of the scattered light and provides a fast and accurate readout of trapped particles’ positions. Using a QPD-based detector scheme exhibiting bandwidth in the MHz-range and calibration methods based on thermal noise analysis of the trapped beads allows for easy, fully automated force calibration. Back focal plane detection, on the other hand, requires a precise alignment of the detector and a careful calibration to map detector voltages to particle displacement.
An interdisciplinary tool
Optical tweezers have found application in biological, medical, physical and chemical investigations, as well as in their interdisciplinary overlap of research. Major advances are being made in the understanding of cellular dynamics by quantifying intracellular forces on a single-molecule level. Also being studied are the mechanical properties and chemically determined force and energy landscapes that regulate single protein folding or the coiling and looping of DNA molecules. Many interdisciplinary research groups focus on the problems in nanomedicine, such as the interaction of the ubiquitous nanoparticles found in many modern products with biological organisms. Being a noninvasive, sensitive force measurement tool with extraordinary sensitivity on the single molecules, optical tweezers have become a reliable tool in the toolbox of a modern interdisciplinary scientist.
Meet the authors
Vitaliy Oliynyk, Ph.D., is a senior applications scientist with JPK Instruments AG in Berlin; email: [email protected]. Philipp Rauch, Ph.D., is a senior applications scientist with JPK Instruments AG in Berlin; email: [email protected]. Stefan B. Kaemmer, Ph.D., is the general manager with JPK Instruments Inc. in Carpinteria, Calif.; email: [email protected].
References
1. A. Ashkin and J.M. Dziedzic. (1987). Optical trapping and manipulation of viruses and bacteria. Science, Vol. 235, pp. 1517-1520.
2. C.L. Asbury et al. (2003). Kinesin moves by an asymmetric hand-over-hand mechanism. Science, Vol. 302, pp. 2130-2134.
3. M.H. Larson et al. (2011). Single-molecule studies of RNA polymerase: One singular sensation, every little step it takes. Mol Cell, Vol. 41, pp. 249-262.
4. J.D. Wen et al. (2008). Following translation by single ribosomes one codon at a time. Nature, Vol. 452, pp. 598-603.
5. J.T. Finer et al. (1994). Single myosin molecule mechanics: piconewton forces and nanometre steps. Nature, Vol. 368, pp. 113-119.
6. H. Sakakibara et al. (1999). Inner-arm dynein c of Chlamydomonas flagella is a single-headed processive motor. Nature, Vol. 400, pp. 586-590.
7. Z. Lansky et al. (2015). Diffusible crosslinkers generate directed forces in micro-tubule networks. Cell, Vol. 160,
pp. 1159-1168.
8. P. Gross et al. (2011). Quantifying how DNA stretches, melts and changes twist under tension. Nat Phys, Vol. 7, pp. 731–736.
9. N. Laurens et al. (2011). Alba shapes the archaeal genome using a delicate balance of bridging and stiffening the DNA. Nat Commun, Vol. 3, pp. 1328-1335.
10. C. Sieben et al. (2012). Influenza virus binds its host cell using multiple dynamic interactions. PNAS, Vol. 109, pp. 13626-13631.
11. D.T. Tambe et al. (2011). Collective cell guidance by cooperative intercellular forces. Nat Mater, Vol. 10, pp. 469-475.
12. R.D. Gonzalez-Cruz et al. (2012). Cellular mechanical properties reflect the differentiation potential of adipose-derived mesenchymal stem cells. PNAS, Vol. 109, pp. 1523-1529.
13. T. Yuan et al. (2015). Diacylglycerol Guides the Hopping of Clathrin-Coated Pits along Microtubules for Exo-Endocytosis Coupling. Dev Cell, Vol. 35, pp. 1-11.
14. H. Wang et al. (2016). Aspect ratios of gold nanoshell capsules mediated melanoma ablation by synergistic photothermal therapy and chemotherapy. Nanomedicine NBM, Vol. 12, pp. 439-448.