Thomas Mayerhöfer,1 Christoph Krafft,1 Ute Neugebauer1,2 and Jürgen Popp1,2,3
1Leibniz Institute of Photonic Technologies
2Center for Sepsis Control and Care, University Hospital Jena
3Institute of Physical Chemistry, Friedrich Schiller University
Raman spectroscopy’s molecular sensitivity makes it promising for clinical applications: It can identify pathogens much faster than current methods, investigate circulating tumor cells, help surgeons distinguish tumors from healthy tissue and identify the chemical nature of cardiovascular plaques and evaluate their severity.
Today’s aging population presents a great challenge: ensuring affordable and sustainable health care systems. As the number of people over 65 grows, society faces an increase in age-related diseases. Accordingly, the number of cases of macular degeneration in Germany, for example, will more than double in the next 15 years. Similar increases have been projected for the number of heart attacks and new cases of dementia, while the number of newly discovered cancer cases is predicted to grow by “only” 50 percent.1 In addition, new problems such as the increasing risk of pandemics will be looking for solutions. At the same time, the often careless use of antibiotics is gradually leading to the emergence of resistant pathogens.2
Meeting all these challenges will require new methods and devices with which – ideally – diseases can be caught and fought before they are even spread.
Photonic methods may be the key. Photonics will allow an understanding of disease development at the molecular level and thus will eventually facilitate a personalized medicine. A particularly promising and versatile photonic method is Raman spectroscopy and its variants. As with all light-based methods, Raman spectroscopy allows noncontact measurement, but unlike fluorescence spectroscopy, for example, it does not require exogenous labels. This means that the technique has the potential to significantly accelerate or even replace laboratory analysis and enable near-patient diagnosis, as it is relatively fast and also very precise. Especially in the field of imaging, the high specificity and low invasiveness must be emphasized. Further advantages of Raman spectroscopy include high spatial resolution, the lack of need for time-consuming sample preparation, and the ability to work in an aqueous environment.
Pathogen diagnosis
Classically, pathogen diagnosis is based on the initiation, growth and analysis of a pathogen culture, a process that can take up to a week and requires experienced personnel. But in the case of sepsis shock, for example, the survival rate decreases dramatically with every hour that passes before targeted treatment – falling to a rate of less than 20 percent after only 12 hours. In the ideal case, an infection would be associated with a pathogen within a few hours.
Because each bacterial species has a unique Raman signature, Raman spectroscopy is well suited to identifying it. In addition, with Raman microscopy, a spectrum of a single bacterium is generally sufficient for identification.3 However, the distinctions between spectra of various species are often subtle, and differentiation by visual inspection is generally not possible. The bacterial spectrum ultimately is the sum of the spectral signatures of all contained substances, such as water, proteins, fats, nucleic acids and carbohydrates (Figure 1a). Even bacteria within the same species show subtle variations caused by differences in growth state, dissimilar conditions among patients such as nutritional status and treatment settings, and varying localization in patients (centers of infection). For pathogens found outside of the patient, environmental factors such as oxygen and carbon dioxide content in the air, temperature and light play a role.
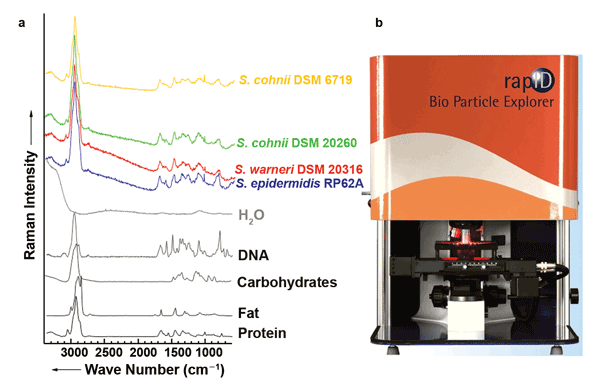 Figure 1. (a) Raman spectra of the main biological components of bacteria: water, protein, nucleic acids (DNA), carbohydrates and fats. In the Raman spectra of various staphylococcal strains that serve as examples, several bands of the individual components can be recognized. (b) The Bio Particle Explorer from rapID. Images courtesy of the authors.
Figure 1. (a) Raman spectra of the main biological components of bacteria: water, protein, nucleic acids (DNA), carbohydrates and fats. In the Raman spectra of various staphylococcal strains that serve as examples, several bands of the individual components can be recognized. (b) The Bio Particle Explorer from rapID. Images courtesy of the authors.
The solution is to apply chemometric techniques to the Raman spectra of bacteria. In this case, a spectrum is broken down into distinct areas that are mathematically compared with equivalent areas in bacterial spectra that have been collected in an extensive database. On average, it is possible to determine almost 99 percent of the bacterial species correctly, and even the strain can be determined with an average accuracy of more than 92 percent.3 To determine bacterial contamination in cleanrooms or air conditioning systems, an appropriate solution is already commercially available (the Bio Particle Explorer from rapID, Figure 1b). This system uses fluorescence spectroscopy to distinguish between inanimate particles and bacteria; the bacteria are then identified by means of Raman spectroscopy.
Its potential for clinical use necessitates its ability to identify bacteria in complex media such as saliva, urine or even blood. In general, bacteria must be separated from these media; otherwise, the medium hampers identification or even makes it impossible. For this separation step, microfluidic chips are currently being developed that use, for example, dielectrophoresis to trap bacteria and make them accessible to measurement.4 This method can also be used to measure bacteria directly in solution and thus permits – in addition to identification – statements about the susceptibility or resistance to antibiotics.
Oncological diagnosis
From carcinogenic tissue, tumor cells can peel off, enter the bloodstream and eventually cause metastases. These isolated tumor cells are relatively accessible and have great diagnostic value. On the one hand, their detection helps to verify the presence of a tumor and to locate and identify it. On the other hand, the stage of the primary tumor can be determined on the basis of circulating tumor cells, and the success of chemotherapy can be assessed.
The principle of detection corresponds substantially to that of optical flow cytometry: The blood is passed through a microfluidic chip; individual cells are then captured by optical traps, examined using Raman spectroscopy, and classified and sorted for further use.
Compared with optical flow cytometry, the Raman method enables a much more accurate diagnosis of the individual cell. The disadvantage is a much lower throughput (five to six cells per minute), but this can be improved upon significantly in the future with device- and component-related upgrades.
Figure 2 shows a microfluidic chip made of quartz for Raman-activated cell sorting.5 The unprocessed Raman spectra are influenced by the spectral properties of the filters, the trapping lasers and the substrate material. Therefore, successful classification requires that these properties be suppressed, so that the spectral fingerprint of white blood cells (green) and tumor cells (orange, brown, blue) is made visible.
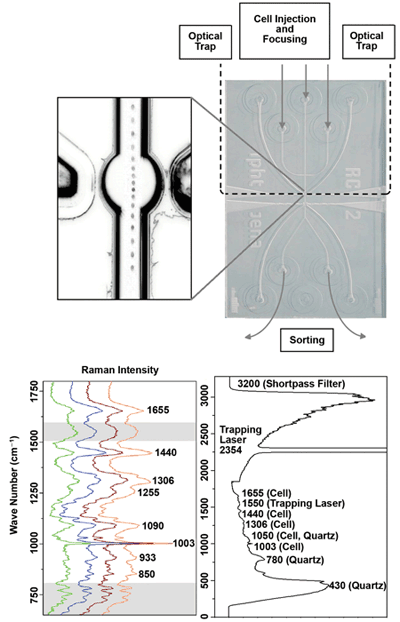
Figure 2. Microfluidic chip made out of quartz for Raman-activated cell sorting.
Radical removal of tumor tissue is not advisable in every case. With brain tumors especially, as little healthy tissue as possible should be removed. Unfortunately, the boundaries between tumor and normal tissues are often diffuse and not clearly defined. To date, tissue removed in the operating room is assessed by a pathologist, who determines whether the tumor has been completely removed. It would be better, however, to have a surgical microscope that could depict the boundaries of the tumor directly. This could be realized by combining several imaging techniques, preferably label-free ones that could enhance the contrast. Particularly promising would be the combination of Raman with coherent anti-Stokes Raman scattering (CARS), two-photon excitation fluorescence (TPEF) using endogenous markers, and second-harmonic generation (SHG).6 While all the (Raman-active) vibration modes are excited simultaneously by Raman scattering, the interaction of three different light pulses isolates a selected vibration that is coherently excited and, in the case of CARS, generates a fourth, spatially oriented and coherent light pulse.
The advantage of CARS over Raman lies in a much shorter image recording time (up to a factor of 104) due to a strong increase in the scattering cross section. In contrast to Raman and CARS, SHG and TPEF emphasize morphological details: SHG is particularly sensitive for ordered noncentrosymmetric structures, such as collagen, whereas TPEF is responsive to endogenous fluorescent substances such as NAD(P)H, flavins and elastin.
Figure 3 compares TPEF, CARS and Raman microscopic images of an unstained thin section of brain tumor with a microscopic image of the same sample after staining with hematoxylin and eosin. The cell nuclei in particular, which are resolved with all methods, are of interest for a histopathologic evaluation. With the help of the combined morphological and functional information through the multimodal approach, chances are good of developing photonic tools that not only can discover and classify tumors early but also allow location of tumor boundaries during surgery.
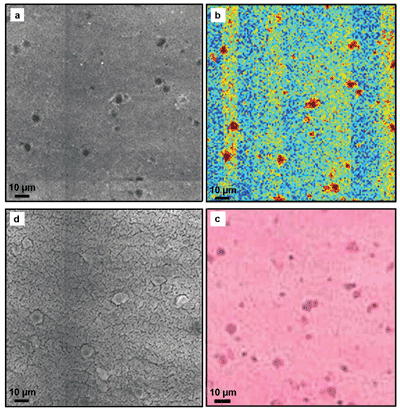

Figure 3. Comparison of (a) TPEF, (b) Raman and (d) CARS microscopy of an unstained thin section of brain tumor with a light microscopic image (c) of the same sample after staining with hematoxylin and eosin.
Cardiovascular disease diagnosis
A potential in vivo application of Raman spectroscopy is the endoscopic examination of arterial plaques. Conventional methods such as intravascular ultrasound or optical coherence tomography generally provide only morphological information. To assess whether a deposit within an artery is dangerous – i.e., can become detached from the vessel wall and cause blockages and thereby heart attacks or strokes – an evaluation of the plaque composition is essential.
The Raman spectra of the potential components of plaques – calcium phosphate, connective tissue, triglycerides
and cholesterol – are well distinguishable, so endoscopic Raman spectroscopy can basically determine the composition of plaque, and thus the danger of deposits.
First experiments on rabbits have confirmed the value of this approach. As shown in Figure 4, a 1-mm-diameter probe with a central excitation fiber and 12 detection fibers for ex vivo measurements was employed7 to record the Raman spectra under in vivo conditions. The signals of plaque deposits differ in the intensity and spectral contributions of lipids from the signals of the arterial wall with collagen bands, and from blood with bands of red blood cells. Future developments aim to combine Raman with optical coherence tomography and/or ultrasound to fuse morphological with molecule-specific information. In addition, miniaturization is planned for examining smaller-diameter arteries.
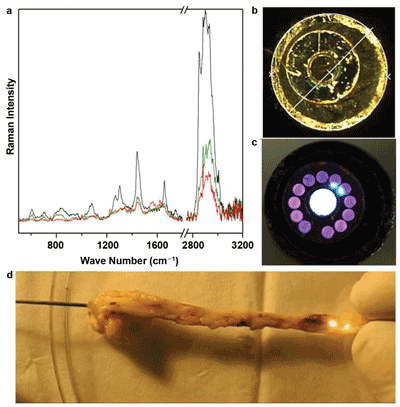
Figure 4. Raman endoscopic examination of the arteries of a rabbit: (a) Raman spectra at different positions inside the artery, (b, c) images of a multicore fiber with 12 collecting fibers around an excitation fiber and (d) recording of the Raman spectra under in vivo conditions.
Meet the authors
Dr. Thomas Mayerhöfer is senior researcher at the Leibniz Institute of Photonic Technologies in Jena, Germany; email: thomas.mayer [email protected]. Dr. Christoph Krafft is head of the Spectroscopy/Imaging work group at the Leibniz Institute of Photonic Technologies; email: [email protected]. Dr. Ute Neugebauer is a junior research group leader at the Leibniz Institute of Photonic Technologies and the Center for Sepsis Control and Care at University Hospital Jena; email: [email protected]. Dr. Jürgen Popp is scientific director of the Leibniz Institute of Photonic Technologies, a member of the Center for Sepsis Control and Care at University Hospital Jena, and director of the Institute of Physical Chemistry at Friedrich Schiller University in Jena; email: [email protected].
Acknowledgment
The authors thank the Thüringer Ministerium für Bildung, Wissenschaft und Kultur (Project-B714-07037), the Federal Ministry of Education and Research (FKZ: 01EO1002/FKZ: 13N10774) and the European Union for financial support.
References
1. F. Beske et al (2009). Morbiditätsprognose 2050: Ausgewählte Krankheiten für Deutschland, Brandenburg und Schleswig-Holstein. Kiel: Institut für Gesundheits System Forschung.
2. European Centre for Disease Prevention and Control (March 2013). Annual Epidemiological Report 2012. Reporting on 2010 surveillance data and 2011 epidemic intelligence data. Stockholm: ECDC.
3. M. Harz et al (February 2009). Vibrational spectroscopy – a powerful tool for the rapid identification of microbial cells at the single-cell level. Cytometry A, Vol. 75A, pp. 104-113.
4. U.-Ch. Schröder et al (2013). Culture-independent detection of sepsis pathogens from bacteriurias: A novel combination of dielectrophoresis and micro-Raman spectroscopy. Infection, Vol. 41 (Suppl. 1), P036.
5. S. Dochow et al (March 2013). Quartz microfluidic chip for tumour cell identification by Raman spectroscopy in combination with optical traps. Anal Bioanal Chem, Vol. 405, pp. 2743-2746.
6. T. Meyer et al (Feb. 10, 2011). Nonlinear microscopy, infrared, and Raman microspectroscopy for brain tumor analysis. J Biomed Opt, Vol. 16, p. 021113.
7. C. Matthäus et al (September 2012). In vivo characterization of atherosclerotic plaque depositions by Raman-probe spectroscopy and in vitro coherent anti-Stokes Raman scattering microscopic imaging on a rabbit model. Anal Chem, Vol. 84, pp. 7845-7851.