Technique combines advantages of atomic force microscopy and Raman spectroscopy.
Dr. Tanja Deckert-Gaudig, Elena Bailo and Dr. Volker Deckert, ISAS — Institute for Analytical Sciences
A fundamental goal in the life sciences is to understand how the elemental building blocks (nucleobases, amino acids, etc.) are composed in living organisms. Chemical processes in cells are carried out by macromolecules such as nucleic acid chains, carbohydrates and proteins. To better understand the biochemical function of cellular components, it is essential to resolve their design. To do so, biochemists need an analytical tool that allows access to structural information in a high-throughput manner.
Tip-enhanced Raman spectroscopy offers a way to elucidate molecule arrangements with a spatial resolution down to 5 to 20 nm. The technique combines the high lateral resolution of an atomic force microscope (AFM) with the chemical information provided by Raman spectroscopy.
Raman spectroscopy can reveal structural details of a sample without labeling. However, in contrast to well-established methods such as fluorescence microscopy, the sensitivity is rather low. To overcome this drawback, tip-enhanced Raman spectroscopy uses a single silver particle attached to an AFM tip to scan the sample of interest while Raman spectra are being simultaneously recorded (Figure 1).1
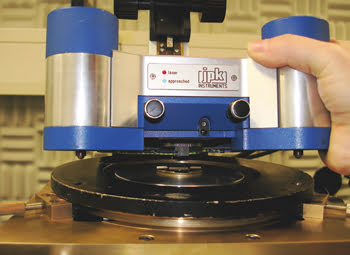

Figure 1. The tip-enhanced Raman spectroscopy setup consists of an AFM and an inverted Raman microscope.
The nanoparticle — which has a diameter of ∼10 nm — leads to a signal enhancement on the order of 106 to 108 and provides a spatial resolution directly related to its size. Hence, Raman spectra can be acquired very quickly. The technique also has the advantages of small sample volumes (<1 μl), short acquisition times and, most important for biological samples, detection at physiological concentrations without further sample preparation.
Jürgen Popp’s group at the University of Jena, together with Volker Deckert’s group at the Institute for Analytical Sciences in Dortmund, both in Germany, have used tip-enhanced Raman spectroscopy on bacterial surfaces.2 The researchers investigated Staphylococcus epidermidis without using labels. This bacterial strain usually is harmless, but it can be pathogenic to people with environmental sensitivities. It binds to polymer surfaces and forms biofilms.
The recorded tip-enhanced Raman spectroscopy bands were identified as peptide and polysaccharide moieties, which agreed with the known chemical composition of the bacteria’s surface. The high-spatial-resolution technique and the few nanometers of penetration depth restricted the spectral measurements to the outermost surface layer of the cells. Thus, the experiments could probe the important class of membrane proteins specifically. These results could lead to a better understanding of biofilm formation.
From surfaces to strands
The highly sensitive technique has great potential for the investigation of dynamical processes on cell surfaces. Because this is an important interface for the cell-cell or cell-drug interaction, the label-free structural investigation of biological processes, specifically at this interface, also could lead to the development of new drugs.
At the Institute for Analytical Sciences, Elena Bailo and Deckert recently applied tip-enhanced Raman spectroscopy to the detection of spatially resolved Raman spectra along a polycytosine RNA strand.3 So far, all known DNA sequencing methods require considerable amounts of DNA and are not in the position for a direct fragmenting of the bases. Therefore, these experiments could provide a further possibility to perform the sequencing in a direct and label-free way.
The spectra in this particular experiment were measured with a spatial resolution down to a few nucleobases (Figure 2). The height of the strand was determined to be ∼1 nm, which agrees with the reported values. The vibrational spectra showed all the spectral features of cytosine. The results of the polycytosine measurements revealed that distinguishing it from other compounds should be possible.
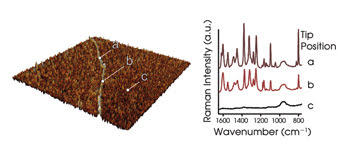
Figure 2. The AFM topography of a single polycytosine (poly-C) strand is shown on the left, and the corresponding tip-enhanced Raman spectroscopy (TERS) spectra are shown on the right. TERS spectra can be detected only when the tip is on top of the RNA.
For a complete target sequencing, it is not necessary to resolve single bases. A sequencing experiment can be performed by moving the tip from base to base along the strand. The spectral changes from point to point then can be assigned to differences in the sequence. Sequencing of RNA strands should be possible, and this principle could be used in a similar fashion to elucidate the structure of all kinds of macromolecular chains.
References
1. R.M. Stöckle et al (Feb. 18, 2000). Nanoscale chemical analysis by tip-enhanced Raman spectroscopy. CHEM PHYS LETT, pp. 131-136.
2. U. Neugebauer et al (Jan. 8, 2007). Towards a detailed understanding of bacterial metabolism — spectroscopic characterization of Staphylococcus epidermidis. CHEM PHYS CHEM, pp. 124-137.
3. E. Bailo and V. Deckert (Feb. 15, 2008). Tip-enhanced Raman spectroscopy of single RNA strands: Towards a novel direct-sequencing method. ANGE CHEM INT ED, pp. 1658-1661.
Meet the authors
Volker Deckert is director of the proteomics department at ISAS — Institute for Analytical Sciences in Dortmund, Germany; e-mail: [email protected].
Tanja Deckert-Gaudig and Elena Bailo are researchers at ISAS.