
Finding particles with machine learning
Hank Hogan
Optical live-cell imaging, in some ways, is a victim of its own success. Because of advances, researchers can collect far more data than they can analyze manually. This is particularly true when it comes to tracking the movement of particles and molecules within a cell. Largely because of problems with particle identification, automated tracking has not worked. Now, particle detection is possible through the use of machine learning methods that distinguish particles by means of characteristic features, according to researchers from Tsinghua University in Beijing, and from Harvard Medical School and Brigham and Women’s Hospital, both in Boston.
The technique eventually could form the basis for automated particle tracking and could be used to study cellular dynamics and to answer basic questions about the transport of molecules within cells, among other things.
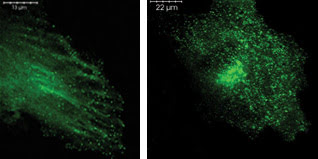
In these images of BSC1 cells, clathrin pits that transport biochemicals are labeled with GFP (left) and a yellow fluorescent protein (right). Researchers developed an automated way to identify such molecular particles, transforming what had been a manual task into one that a computer could handle. The computerized technique could eventually form the basis for automated tracking of the dynamic behavior of clathrin pits and other molecular particles. Courtesy of Stephen T.C. Wong and Xiaobo Zhou, Harvard Medical School.
Thanks to the advent of digital imaging technology, researchers regularly capture high-content images that may contain the location of thousands of moving particles or molecules in a single cell. But many of these images have low contrast and a noisy background, and the particles are often only a few pixels across and have edges that are indistinct. In an image, the particle may appear more like a shapeless blob than a sharp point.
The result is that traditional automated detection methods don’t work. The lack of a sharp edge makes it difficult to distinguish the particle from the background, and the lack of contrast makes it hard to get an accurate automatic picture of the number of particles present. Thus, researchers have had to resort to time-consuming manual methods to extract such particle data.
The team applied machine learning to the problem, teaching computer algorithms how to distinguish a particle from what is not a particle through training data. It chose to use Haar features to make the distinction. Haar techniques were developed in part to ease the computational burden arising from image analysis using data such as pixel intensity alone. Haar methods look at rectangular regions in an image and sum up the number of pixels per region. The resulting value then is used to classify the image and to categorize an area as either having a particle or not, for example.
In the case of particles within a cell, Haar features are a combination of the intensity, shape and size of the objects. To get around fluorescence intensity fluctuations in classifying objects, the researchers relied upon signal-to-noise ratios to decide which automatically identified particles were valid. They discarded those with too low a ratio and kept those with a ratio above a threshold.
They applied these techniques to cell images, looking for clathrin-coated pits that are involved in moving proteins and lipids from the plasma membrane to elsewhere within the cell. They first acquired images with a spinning disk confocal head from PerkinElmer of Boston coupled to fully motorized epifluorescence microscopes from Carl Zeiss of Thornwood, N.Y., with a cooled CCD camera from Photometrics of Trenton, N.J. They imaged using total internal reflection fluorescence and fluorescence microscopy and performed analysis with the public domain software OpenCV, which has been successfully used for human face detection.
They next constructed a training set consisting of 10 × 10-pixel subwindows. Some were selected because they did not contain any particles of interest — a negative sample. Others were selected as positive samples — images that did contain particles. Using the training set, they developed a classifier through machine learning, employing cascading tests in which potential particles had to survive multiple assessments before finally being categorized as a particle.
As a final step, they applied a signal-to-noise ratio threshold to the data to separate particles from nonparticles. After the particles had been detected, they processed the data further to extract the boundary and area of the particles, information needed for accurate tracking.
With this approach, the researchers looked at the data captured in various movies, one based on total internal reflection fluorescence, another on fluorescence and two more on other methods. They compared the results from the automated analysis with that of a manual approach to determine the true positive and false-positive rates. They found that the true positive rate averaged more than 98 percent. Thus, more than 98 out of 100 times, the algorithm successfully identified a particle as a particle. The false-positive rate was 4.44 percent, indicating that most of the time what the algorithm identified as a nonparticle was indeed that. The work is detailed in the August issue of Cytometry Part A.
The researchers noted that the technique could provide a cost-effective automated solution to detecting particles within living cells. The next step, they reported, will be to use the information extracted with this method to track molecular particles. That work is ongoing.
Contact: Stephen T.C. Wong, HCNR-Center for Bioinformatics, Harvard Medical School and Brigham and Women’s Hospital, Boston; [email protected].
Published: September 2007