An algorithm classifies pathology samples as benign or malignant.
Hank Hogan, Contributing Editor
Prostate cancer, according to the American Cancer Society, is among the most common malignancies in men and is the second leading cause of death, accounting for about 10 percent of male cancer-related mortality. If the disease is caught early, the chance for survival is nearly 100 percent. If it is diagnosed after having spread to distant areas, the five-year survival rate is one in three.
It is important to separate the good from the bad so that malignant tissue can be properly treated. Computerized image analysis should be of great help in classifying pathology samples as benign or malignant. However, automatically determining malignancy has proved difficult.
Now researchers from the University of Tehran in Iran and from the Henry Ford Health System and from Wayne State University, both in Detroit, have demonstrated an image analysis approach that automatically spots prostate cancer. The technique has achieved accuracy rates as high as 98 percent in classifying tissue seen in prostate pathology images as benign or malignant.
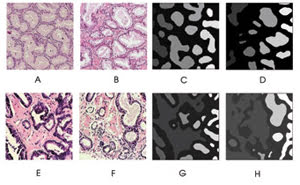
Automated image analysis can detect prostate cancer. These examples show prostate biopsy images under various magnifications and illuminations. Benign samples (A,B) have glands of about the same size that are equally spaced, whereas malignant specimens (E,F) have fused, irregularly shaped glands. After undergoing processing that segments glandular regions, the malignant samples (G,H) display roughly triple the normalized variance in gland size and roundness compared with benign samples (C,D); these criteria can be used to discriminate between the two types of tissue. Reprinted with permission of Cytometry Part B: Clinical Cytometry.
Knowing the degree of malignancy is important when planning treatment; that assessment is currently made by expert pathologists. Unfortunately, this determination is subjective, with pathologists making conflicting diagnoses. And this manual approach takes time and is somewhat difficult.
An automated pathologist
To devise an approach to automate the process, the investigators attacked the problem by imitating how pathologists establish the degree of malignancy. They examine the structure of glands, which consist of lumina, stroma and nuclei. In normal tissue, the glands are a round mass arranged uniformly and are all approximately the same size. In cancerous tissue, the glands are fused and in irregular patterns. Pathologists grade the tissue based on how well it fits these criteria.
The algorithm developed by the investigators first converts the image to gray scale, removing color variations that arise from staining differences. Further processing enhances the nuclei, and then roughness information allows the exclusion of nuclei from stroma and lumina. From this data, the researchers extract glandular features. Numbers derived from variance and roundness are used to distinguish between benign and malignant tissue, with the formulas set up by the researchers such that higher numbers correspond to a greater proportion of malignant tissue. The two values can be combined into one overall cancer index; values above a threshold indicate malignancy.
As detailed in the July issue of Cytometry Part B: Clinical Cytometry, the scientists applied the technique to a dataset of 91 images and one of 199 images. The first set consisted of images of similar magnifications and illuminations, whereas the second had differences in both. Each set contained images from tissue classified by pathologists as completely benign at a Gleason grade of 1 or 2, clearly malignant at a 4 or 5, and possibly normal but probably malignant at a score of 3. On the first set of data, the algorithm correctly classified the samples 97.8 percent of the time, while on the second set it was right 95.5 percent of the time. On the whole, in terms of glandular tissue, the lack of roundness proved a better cancer determinant than variation in size.