Researchers are developing fluorophores and probes that are brighter and that can improve various biological studies.
Hank Hogan, Contributing Editor
Fluorescent probes are small but powerful, shedding light on the inner working of cells and the diseases that afflict them. However, sometimes the right probe for a particular job does not exist. That is why research and development of new probes continues. A look at recent efforts from around the globe finds new probes that track glucose usage, that are brighter by design, that highlight dying cells and that pack dyes together without having them get in each other’s way, fluorescently speaking.
Researchers from Seoul National University in South Korea have developed a fluorescent glucose bioprobe that might someday benefit the millions worldwide who suffer from type II diabetes. It is unlike others in that it operates with the biologically important form of the sugar D-glucose present.
“Researchers now can monitor the glucose trafficking under physiological conditions in the presence of D-glucose in real time using fluorescence imaging equipment without any biohazards,” said Seung Bum Park, a chemistry professor at the university and leader of the research group.
Because glucose is an important source of cellular energy, assessing its use can yield vital diagnostic information. Unfortunately, existing probes suffer from various drawbacks. One such probe —dubbed 18FDG for its isotopic and chemical makeup — is used for tumor diagnosis via PET, but it is short-lived and a biohazard. A fluorescent bioprobe, named 2-NBDG in part because it attaches a fluorophore to the molecule through a nitrogen atom, eliminates these problems. However, it works only in sugar-depleted environments and so is not suitable for living cells under normal conditions.
To synthesize a new bioprobe, the researchers took a different approach. They attached a dye to the D-glucose, or dextrose, molecule. After an initial approach failed, they changed the synthesis and optimized it to yield a stereochemically pure compound to which various fluorophores and linkers could be introduced. For their initial demonstration, they used the fluorescent dye Cy3 because it resists photobleaching and is compatible with various bioassays.
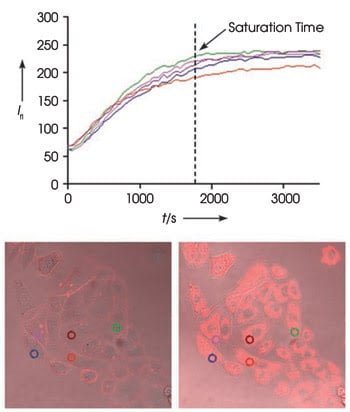
The uptake of a fluorescent glucose bioprobe by A549 cancer cells (top) drives fluorescence intensity (Ifl ) up as measured at the five locations indicated in the circles in the bottom. On the bottom are merged phase contrast and fluorescence images. The left images show the initial view; the right, after 60 min. Reprinted with permission of Angewandte Chemie International Edition.
After determining that the bioprobe followed the uptake pathway of D-glucose, the researchers tested it by looking at the effect of an anticancer agent on tumor cells. “The enhanced glycolysis of cancer cells should have a significant reduction of glucose uptake in cells treated with anticancer agent,” Park explained.
Using Zeiss inverted and confocal laser scanning microscopes, they measured the uptake of the bioprobe 3, 6, 12 and 24 h after treating human A549 cancer cells with various concentrations of taxol, averaging the fluorescence intensity from 35 to 50 cells in each reading. The uptake fell by nearly 50 percent 6 h after treatment with a 9.8 μM taxol concentration but <5 percent with a 49-nM dose.
Park predicts that the bioprobe will find use in studies involving metabolic syndrome, a condition characterized by diabetes and obesity. The probe could be used to screen small molecules that modulate glucose uptake, something that could lead to therapeutic agents. Such research is hard to do today, Park said. “There are quite limited research tools for high throughput/content screening for metabolic disease.”
The work is detailed in the March 12 issue of Angewandte Chemie International Edition.
Getting it brighter
By knowing not only when but also how to fold them, researchers at the University of Amsterdam and at Utrecht University, both in the Netherlands, have produced improved green and blue fluorescent proteins.
These strongly enhanced green fluorescent (SGFP2) and strongly enhanced blue fluorescent (SBFP2) proteins are several times more photostable and have a quantum yield several times that of their predecessors. These characteristics mean that more photons can be detected for a prolonged time, said Theodorus W.J. Gadella Jr., a molecular cytology professor at the University of Amsterdam.
He added that folding and monomerization also are important because improper folding and molecules that clump decrease fluorescence intensity and cause unwanted interactions in cells. Tests show that the new fluorescent proteins are much brighter when expressed by living cells, particularly in bacteria.
The researchers began with the Venus yellow fluorescent protein — the brightest and most efficiently folding variant known. They mutated the fluorescent protein by inducing changes through site-directed mutagenesis that produced fluorescent proteins of different colors. They tweaked it to further improve performance.
They measured the performance of both, characterizing SGFP2 and SBFP2 with a Bio-Tek Instruments dual-beam spectrophotometer and a Photon Technology International fluorescence spectrofluorometer. When determining photobleaching, they used a Zeiss inverted microscope to illuminate the proteins and a Roper Scientific CCD camera, along with various filters, to record the fluorescence decay. Another characterization tool they used was fluorescence lifetime imaging in a time-correlated single photon counting setup with a Nikon microscope, PicoQuant pulsed picosecond diode lasers, and a Hamamatsu photomultiplier tube. Finally, they performed confocal microscopy to image cells, using a Zeiss confocal laser scanning microscope.
As described in the Mar. 27 issue of Biochemistry, the absorbance and emission peaks of the new proteins were within a few nanometers of earlier versions. However, the intrinsic brightness of SBFP2 was 3.5 times and its photostability double that of enhanced BFP.
Bacteria expressing SBFP2 were 16 times as fluorescent as those expressing enhanced BFP, and those expressing SGFP2 were seven times as bright as those expressing enhanced GFP. In mammalian cells, the ratios were 3.7 and 1.7 for SBFP2 and SGFP2 as compared with enhanced BFP and enhanced GFP, respectively.
The huge increase observed in bacteria was a result of their lack of specific molecules that chaperone the folding of the proteins, according to Gadella. “In bacteria, the efficiency of self-folding is a major determinant for brightness. In mammalian cells, folding is assisted by chaperones, and here the brightness is more determined by the intrinsic brightness.”
The scientists are working to further increase the fluorescence quantum yield. Although SGFP2 and SBFP2 are the third or fourth generation of fluorescent proteins derived from the original GFP, the researchers feel that there is still room for further improvement.
Tracking cell death
One way that cells die is by an orderly process called apoptosis. Because apoptosis plays a role in organism development, self-regulation and treatment response, scientists would like to non-invasively monitor the process in living cells and in whole animals. Now researchers from the Washington University Medical School in St. Louis have created a fluorescent probe for this purpose.
The probe works well for tracking apoptosis for several reasons. It is cell-permeable and activated by caspase 3, one of the protease effectors of apoptosis. Unlike initiator caspases, the presence of an effector indicates that cell death is under way. Most fluorescent probes do not target caspase 3 at all. Antibodies are specific to the biomolecule, but they require fixing and processing cells, procedures not compatible with in vivo research. In addition, other fluorescent probes emit in the blue-green range, which is not optimal for in vivo applications. This probe emits in the near-infrared.
At the heart of the new probe is an old idea, a quenching-activation strategy. The probe consists of an effector caspase recognition sequence flanked by a fluorophore and a complementary quencher. When the caspase cleaves the construct in twain, the quencher is separated from the fluorophore, and what had been a dimly fluorescent probe suddenly becomes much brighter. The work appears in the April 3 issue of Biochemistry.
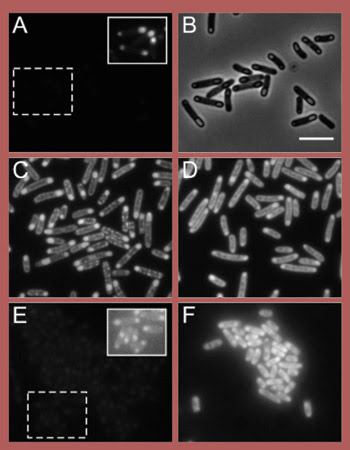
These fluorescence micrographs of E. coli show various green and blue fluorescent proteins (original proteins shown in A and E, respectively) with various induced mutations (C, D and F) designed to improve intrinsic and expressed brightness. B is a phase contrast image of A, and the insets in A and E show contrast-enhanced images of the region of interest. Reprinted with permission of Biochemistry. Scale bar = 2 μm.
Dr. David Piwnica-Worms, a professor of radiology, molecular biology and pharmacology, led the research team. He noted that prior work looking at this approach for activation has focused on large nanoparticle platforms with appended peptides. The resulting structures generally could not be used inside a cell. No one had tried intramolecular quenching strategies with a small molecule attached to a peptide that allowed the cell membrane to be permeated and the interior of the cell reached.
The investigators synthesized their substrate, dubbed TcapQ647, using Alexa Fluor 647 for the fluorophore and QSY 21 as the quencher. Experiments showed that caspase 3 cleaved the substrate much more efficiently than either caspase 6 or 7, two other effectors, and that the initiator caspases did not cleave at all.
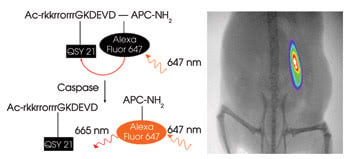
A probe (left) consists of an effector caspase recognition sequence flanked by a fluorophore (Alexa Fluor 647) and a quencher (QSY 21). When exposed to caspase 3, an effector caspase present during cell death, the quencher is separated from the dye, and it becomes fluorescent. On the right is an image of parasite-induced apoptosis of a xenograft in a mouse model. Reprinted with permission of Biochemistry.
Using a fluorescence spectrophotometer from Varian Inc. of Palo Alto, Calif., the researchers illuminated the probe at 647 nm and captured emission at 670 nm, measuring fluorescence before and after cleavage. They also used a Zeiss laser scanning confocal microscope to monitor the results in living cells undergoing chemotherapy-induced apoptosis and an Eastman Kodak fluorescence imaging instrument to detect parasite-induced apoptosis as it progressed in a mouse model.
Piwnica-Worms said that they are working on improving the probe and expanding the platform to other protease activities, among other research avenues. “We are also combining the approach with selected therapeutic strategies,” he said.
Overcoming quenching
With fluorophores, more sometimes can be less effective because multiple tags can suppress fluorescence. Now a group from Carnegie Mellon University in Pittsburgh has created supramolecular fluorescent labels that do not have this problem. These nanostructured DNA templates keep the fluorophores close, but not too close.
Associate professor of chemistry Bruce A. Armitage, one of the team leaders, explained that the concept arose from a discussion of a class of molecules called phycobiliproteins. “Each protein has dozens of chromophores bound to it, but the protein keeps each chromophore sequestered from the others to minimize self-quenching,” he said.
To implement this concept in their studies, the researchers placed fluorophores between the base pairs of DNA via intercalation, with a maximum of one dye every other base pair. That produced a minimum separation of ~7 Å. A similar concept using linear DNA had been described and patented a decade ago by Alexander Glazer at the University of California, Berkeley.
Armitage and his colleagues used both linear and branched DNA nanostructures, dubbed “nanotags,” with the most common being three 10-base-pair arms joined at a junction. Thus, up to 15 chromophores could be bound to each nanostructure.
For experiments, they used a yellow chromophore noncovalently bound to the DNA nanostructure. They found steadily increasing fluorescence intensity compared with chromophore added to a solution containing the nanostructures until a ratio of one chromophore to two base pairs was reached. This situation was in marked contrast to the usual tendency of organic dyes in solution to clump and self-quench.
The DNA nanostructure also was investigated for effectiveness in Förster resonance energy transfer (FRET) experiments, in which a fluorescent donor absorbs light and then transfers the energy to an acceptor.
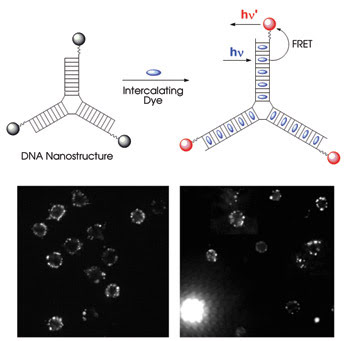
With the appropriate scaffold, fluorescent dye molecules can be packed together and not self-quench. In the top diagram, a three-way junction DNA nanostructure holds intercalating fluorescent dyes far enough apart so that they do not interfere with each other’s fluorescence. With the correct acceptor attached to the end of the structure, the intercalated dyes act as donors and so exhibit very efficient FRET. The nanostructures work as tags (bottom), as shown in these fluorescence microscope images comparing mouse T cells labeled with traditional reagents (left) and the new nanostructures (right). The bright structure in the bottom left of the nanostructure image is a dead cell, highlighted because the intercalated dye dissociated from the nanostructure. Reprinted with permission of the Journal of the American Chemical Society.
In one experiment, the scientists attached Cy3 acceptors to the end of the nanostructure and demonstrated FRET efficiencies of 95 percent. The structures harvested light at 440 nm and emitted light at 570 nm. The work was published in the Feb. 21 issue of the Journal of the American Chemical Society.
In a proof of concept, the researchers performed flow cytometry on polystyrene microspheres coated with streptavidin. They combined these with nanostructures functionalized with biotin, a molecule with a high binding affinity for streptavidin. They also successfully labeled mouse T cells, achieving staining patterns that were similar to those produced by conventional fluorescent immunophenotyping reagents.
Although promising, aspects of this DNA nanostructure approach need improvement. Because of the noncovalent binding, the dye can dissociate from the nanostructure, diminishing the signal and potentially binding to other molecules. “We are working very hard on generating nanotags where the dyes are covalently attached to the DNA to prevent these problems,” Armitage said.
He added that they are looking into using three-dimensional DNA structures and not just two-dimensional ones. The ultimate goal is to use the nanotags to image individual molecules in cells but, Armitage noted, challenging obstacles remain.