From cancer diagnosis and prognosis to the effects of applying medical leeches, the applications of Maldi imaging are growing.
Higher throughput and a wider choice of instruments such as solid-state UV lasers have greatly improved the analytical capabilities of matrix-assisted laser desorption/ionization (Maldi) mass spectrometry (MS). Maldi-MS is making a strong comeback, competing with many applications classically performed by electrospray ionization, especially in terms of analytical speed.
In particular, new approaches for absolute quantification are performed better and at higher throughput with Maldi-MS. Moreover, recent work to combine Maldi with a range of mass analyzer techniques is expanding its analytical potential.
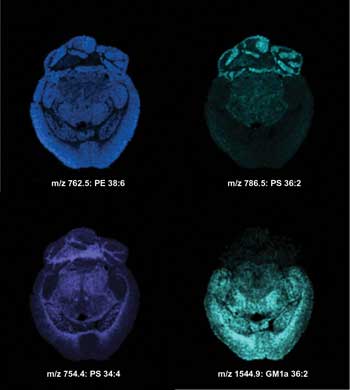
Imaging MS of lipids in the negative ionization mode from a transversal mouse brain section after matrix sublimation. Ion images were acquired with a lateral resolution of 100 µm. m/z = mass-to-charge ratio; PE = phosphatidylethanolamine; PS = phosphatidylserine; GM1a = ganglioside; ST (OH) = sulfatide alcohol. Courtesy of the University of Montreal.
Maldi is one of two existing techniques (along with electrospray ionization) that allow ions to be produced from high-molecular-weight compounds, such as proteins, in an intact manner. These ions can then be analyzed by mass spectrometry.
The method involves mixing an analyte molecule with a matrix, typically in a 1:5000 ratio, said Dr. Pierre Chaurand, associate professor at the University of Montreal. Most of the commonly used matrices are small organic acids, with the essential property of absorbing at the wavelength of the ionizing laser.
“Upon mixture deposition on target and solvent evaporation, matrix-analyte co-crystals form,” he said. “When these crystals are irradiated with brief pulses of UV laser light, the matrix absorbs the light and is instantaneously volatilized. This releases (desorbs) the analyte from the surface and ionizes it.”
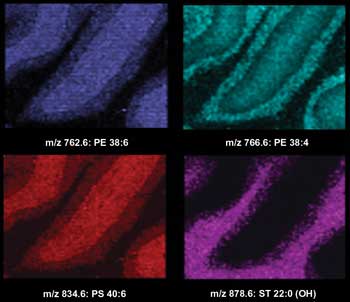

Imaging MS of lipids in the negative ionization mode from a transversal mouse brain section after matrix sublimation. Ion images were acquired with a lateral resolution of 100 µm. Courtesy of the University of Montreal.
When Maldi imaging first emerged in the late 1990s, scientists focused on improving technological performance, such as developing better sample-preparation methods, bioinformatics tools and instruments.
Although Maldi-MS of homogenized samples has been around for a long time, groups such as Chaurand’s are now carrying out Maldi-MS while keeping the tissue intact.
Professor Isabelle Fournier, Maldi imaging team leader at the University of Lille in France and a pioneering scientist in the field, admits that only in the past two to three years have researchers started to discover the range of biological applications.
Almost every kind of biomolecule can be analyzed using Maldi imaging, including proteins, peptides, nucleic acids, lipids and other metabolites. This information can be used to determine tumor margins, drug distribution throughout a tissue or organism, and proteomic profiles under various experimental or therapeutic conditions.
“From these applications, what we can say is that Maldi imaging is useful for defining areas of distinct molecular contents in tissues – i.e., corresponding to different physiological processes,” Fournier said. “In oncology, this is useful for finding cancerous areas and normal areas as well as interfaces where cells are already changing. Therefore, this is a technology that could greatly help pathologists for their diagnosis by bringing the molecular dimension.”
In fact, she has witnessed an increase in the use of this technology in hospital pathologists’ services over the past five years. And one of her aims is to develop a system for in vivo imaging of patients using fiber lasers rather than invasive biopsies.
In recent years, new techniques and instruments have enabled scientists to explore a wider range of applications, including the study of Parkinson’s and Alzheimer’s diseases, tumor diagnosis and prognosis as well as more fundamental studies.
Historically, Maldi ion sources were coupled to time-of-flight mass analyzers. But in the past five years, they have been adapted for use with a wide range of other mass analyzers – quadrupole time-of-flight, ion trap, triple quadrupole, Fourier transform ion cyclotron resonance, etc. – enabling the imaging of drugs, lipids and metabolites from tissue sections with very high mass resolving power and accuracy.
When it comes to Maldi-MS, lasers are a critical and often rate-limiting step in the sampling process. Matrix solutions can be applied in many different ways, such as spraying and spotting. In the latter case, the matrix spot may be around 100 µm and consist of crystals scattered around the spot and formed by the matrix and incorporating the analyte.

Spotting reagents provide flexibility of reagent types, timing control, repeatability and localization of the analysis area to specific areas such as clusters of cancer cells. The Portrait 630 reagent multispotter uses focused acoustic energy to generate droplets of reagents, including Maldi matrix solutions, buffers and surfactants. Acoustic transfer has no nozzles to clog, enabling even the highest concentrations of crystallizing agents. Courtesy of Labcyte Inc.
Irradiating the sampling spot involves many laser shots; ideally, the diameter of the light on this spot and the pulse rate of the laser must be as small and fast as possible to feed the mass spectrometer.
The emergence of UV solid-state lasers at wavelengths of 355 and 349 nm enables Maldi-MS to be performed at kilohertz repetition rates, which greatly improves analytical throughput.
A better understanding of diseases
Chaurand and colleagues at the University of Montreal currently are working on introducing Maldi imaging MS to the clinical setting as a complement to conventional histopathology for disease diagnosis and prognostic purposes.
“We hope to set up a pan-Canadian consortium on imaging MS centered on several imaging-MS-capable academic labsin close partnership with clinical institutions,” Chaurand said. “The goal is that disease-specific molecular patterns will help clinicians better assess the status of the disease and ultimately develop personalized therapy.”
In this pursuit, Maldi-MS is used to investigate the molecular content of tissue sections and directly correlate this molecular information with the histology of the tissue sections that have been observed by optical microscopy.
The molecular profiles and images recovered display unique features that correspond with the cell type locally investigated (October 2007 Nature Methods, doi: 10.1038/nmeth1007-781). This has direct implication for the study of disease. For example, the molecular signatures recovered from tumors are radically unlike those observed from healthy tissues.
Chaurand said that imaging MS is potentially powerful because it could combine molecular specificity with traditional histopathology for diagnostic purposes.
In a Nature Reviews Cancer article published in September 2010 (doi: 10.1038/ nrc2917), Kristina Schwamborn and Richard Caprioli at the Mass Spectrometry Research Center at Vanderbilt University in Nashville, Tenn., put forward Maldi-MS as a new and effective tool for molecular studies of complex biological samples such as tissue sections.
Histological features remain intact throughout the analysis of a section, so distribution maps of multiple analytes can be correlated with histological and clinical features. This means that spatial molecular arrangements can be assessed without the need for target-specific reagents. More importantly, diagnostic and prognostic markers of different cancer types can be discovered, allowing effective therapies to be determined.
Fournier and colleagues have used the technique to screen cancer biopsies and compare them with biopsies of ovarian benign pathologies. This has led the team to identify several potential markers that could be used for early diagnosis of ovarian cancer.
The effects of leeches and drugs
In a more unusual application, the group led by Fournier recently looked at the regeneration processes of the central nervous systems of animals after the application of medical leeches (April 2011 PLoS One, doi: 10.1371/journal.pone. 0018359).
“We have been using Maldi imaging to identify several molecules (lipids, peptides, proteins) that are key players of this biological process,” Fournier said.
But a wider application may be emerging in the field of drug development. Fournier is co-founder of the startup ImaBiotech SA of Lille, France, which proposes the use of Maldi imaging on whole-body animal tissue sections to localize a drug after its administration and to follow its subsequent kinetics.
Pharmaceutical companies such as Novartis Institutes for Biomedical Research of Basel, Switzerland, are starting to take notice of Maldi imaging for this purpose because – unlike existing methods such as autoradiography – Maldi imaging does not require labeling.
It’s all about the preparation
Although the diagnostic and prognostic potential of imaging MS has been clearly established at the protein level, the study of biomolecules such as lipids and metabolites requires new, robust sample preparation methods and protocols.
Preparation of samples is the key focus at Labcyte Inc., a global life sciences instrumentation company with headquarters in Sunnyvale, Calif. In particular, the company specializes in the deposition of a matrix directly onto tissue sections.
“Now the sample can be analyzed without first being homogenized, as is usual with Maldi analysis,” said Richard Ellson, chief technology officer at Labcyte. “By placing a small spot of matrix on the tissue, we can localize the source of the analyte in relationship to the physical structure of the tissue.”
By laying down a regular grid of spots – such as pixels in a color image – an image of the tissue can be built. However, rather than a color image, a pattern of protein expression is constructed.
But it is tricky work to transfer the matrix in this process: The matrix solution is highly volatile and contains a high concentration of dissolved materials.
“Nozzle-based systems will clog – when solvent evaporates, the solute will precipitate,” Ellson said. “Our instruments use the acoustic process we developed that has no nozzle and thus avoids this issue, providing a robust method that can be trusted on one-of-a-kind samples such as tumor biopsies. For tissue-imaging applications, our Portrait reagent spotter targets droplets to the same location to enable chemical process to occur in a localized area (like breaking big proteins into smaller ones) prior to matrix deposition.”
Over the past few years, Labcyte has received much interest from pathologists who are keen to employ this method for understanding and diagnosing cancer. Interest stems from the ability the method offers to select specific locations, such as cancer cell clusters, for analysis.