
Optogenetic Tool Guides Brain Mapping
A new optogenetics technique could help map neural networks in living organisms.
Developed by a team at Caltech, the technique offers a clear view of the brain’s circuitry in action, from the moment a neuron fires to the resulting behavior in the whole organism.
“Our overarching goal for this tool was to achieve sensing of neuronal activity with light rather than traditional electrophysiology,” said Dr. Viviana Gradinaru, an assistant professor of biology and biological engineering at Caltech.
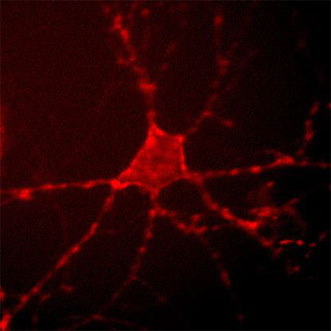
Archer1 fluorescence in a cultured rat hippocampal neuron. Researchers tracked the electrical activity of the cell by monitoring changes in this fluorescence signal at up to a thousand frames per second. Courtesy of Nicholas Flytzanis, Claire Bedbrook and Viviana Gradinaru/Caltech.
The researchers optimized archaerhodopsin (“Arch”), a protein derived from bacteria that acts as a neural inhibitor when exposed to green light.
When it is exposed to red light, the protein acts as a voltage sensor, responding to changes in membrane voltages by producing a flash of light.
These traits could allow Arch to detect and map the activity of neuron networks, but “Arch fluorescence is very dim and is not optimal for applications in live-cell imaging,” the researchers wrote in their study, which was published in Proceedings of the National Academy of Science (doi: 10.1073/pnas.141398711).
They used “directed evolution to identify mutations that dramatically improve the absolute brightness of Arch,” ultimately producing proteins that were 20 times brighter than the original Arch.
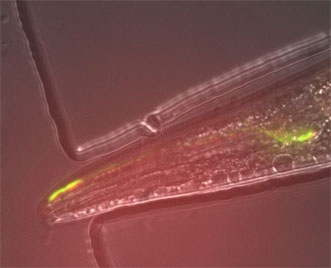
Archer1 fluorescence in an AWC sensory neuron in a live, behaving worm. The response of this cell to stimulus is monitored through tracking changes in fluorescence post-stimulation. Courtesy of Hui Chiu and Viviana Gradinaru/ Caltech.
In a separate, related study, the researchers genetically incorporated the brighter Arch protein variants into rodent neurons to see which version was most sensitive to voltage changes.
One of these variants, Archer1, was not only bright and sensitive enough to mark action potentials in mammalian neurons in real time, it could also be used to identify which neurons were synaptically connected and communicating with one another in a circuit.
“What was interesting is that we would see two cells over here light up, but not this one over there, because the first two are synaptically connected,” Gradinaru said. “This tool gave us a way to observe a network where the perturbation of one cell affects another.”
The researchers also tested Archer1 as a sensor in C. elegans. The researchers found that when a stimulant was present, a baseline fluorescent signal could be seen; when the stimulant was removed, the circuit of neurons lit up. This demonstrated that these neurons were repressed in the presence of the stimulus and active in its absence.
Gradinaru next hopes to use such genetic tools to better understand the complex neuronal networks of mammals, using microbial opsins as sensing and actuating devices in optogenetically modified rodents.
The study was published in Nature Communications (doi: 10.1038/ncomms5894).
Both studies were funded by the National Institutes of Health, the U.S. Army Research Office, a Caltech Biology Division training grant, the Caltech Division of Biology and Biological Engineering, the Shurl and Kay Curci Foundation and the Life Sciences Research Foundation.
For more information, visit www.caltech.edu.
Published: September 2014