On the road to ultrafast imaging, bottlenecks include both hardware and software – but researchers are working to overcome these challenges.
Being able to take a faster snapshot could benefit bio researchers – and the rest of us, too. Faster imaging would allow researchers to capture currently hidden cellular processes, to screen large numbers of cells for relatively rare specimens, and to better reveal how the brain and heart work. The resulting data would be a boost to biology, medicine and related fields.
But faster snapshots need faster cameras, along with other advances in hardware and software. Consider research being done at Stanford University in California, where Dr. Hongjie Dai, a chemistry professor, heads a group pushing to extend ultrafast bioimaging into a new spectral region.
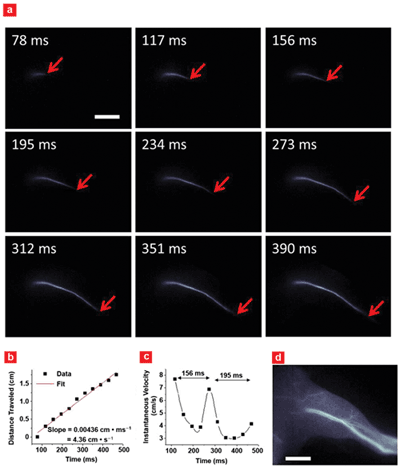
Imaging of arterial blood flow using the second near-IR window, 1000-1700 nm. (a) Blood flow inside mouse femoral artery after fluorophore injection. (b) A plot of blood flow front as a function of time with (c) instantaneous velocity changing due to cardiac cycles. (d) A fluorescence image of the hind limb after fluorophore perfusion. Scale bars in (a) and (d) indicate 5 mm. Courtesy of Hongjie Dai, Stanford University.
“We believe the development of new camera with sensitivity in the second near-infrared window – or from 1000 to 1700 nm – and fast data acquisition rates of a kilohertz and even higher will enable biologists working on cells and live animals to study ultrafast biological and even molecular processes with unprecedented spatial and temporal resolution, as well as deep penetration depths into the animal body,” Dai said.
One benefit of these wavelengths is that there is little autofluorescence and scattering from tissue, allowing imaging with penetration depths of up to a centimeter. In contrast, imaging at shorter wavelengths, such as the visible and near-infrared below 1000 nm, is limited to only a few hundred microns’ depth at best, Dai indicated.
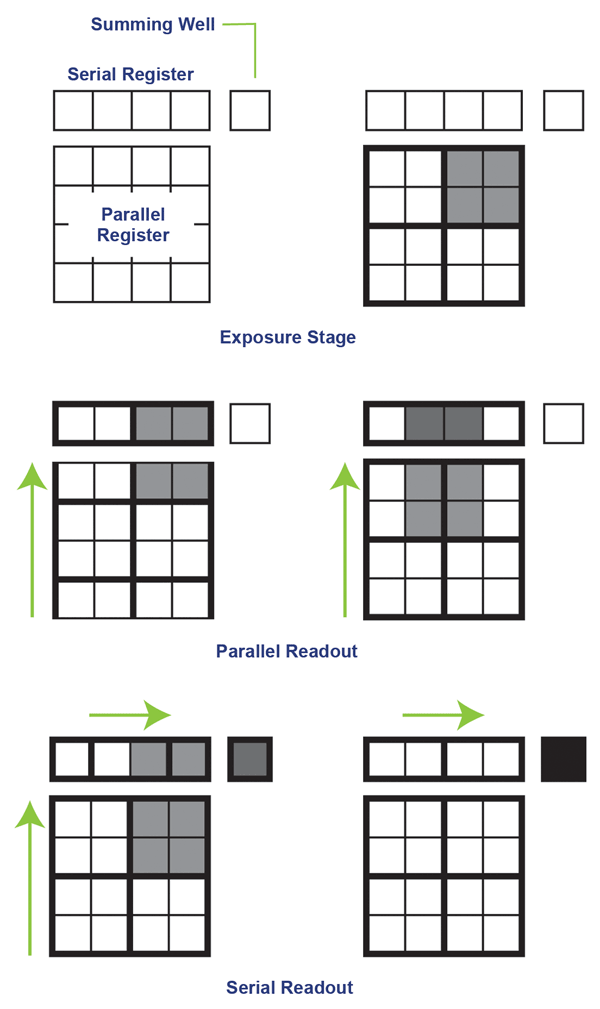 Binning, which combines pixels or only works with a small pixel window, can increase imaging speeds from tens or hundreds to thousands of frames per second. Courtesy of Photometrics.
Binning, which combines pixels or only works with a small pixel window, can increase imaging speeds from tens or hundreds to thousands of frames per second. Courtesy of Photometrics.
With these longer wavelengths, live animals can be imaged – not just cells. One possible application is real-time functional imaging of cardiac cycles in live animals. Dai and others have done this for a single cardiac cycle of a mouse, which takes place over about 200 ms, describing this in June in Nature Communications (doi: 10.1038/ncomms5206). Another application is high-speed imaging of fast dynamic processes at the cellular and subcellular levels, according to Dai.
There are a few hurdles to overcome, however. One is a lack of fluorophores that can be used to label and thus image biological compounds and processes of interest. Dai’s group is working on this, with progress reported in several papers over the past few years. Some of the latest research has used conjugated polymer fluorophores with tunable emission wavelengths running from 1050 to 1350 nm. Other possibilities are fluorophores based on carbon nanotubes or quantum dots.
A second challenge is a lack of cameras. Silicon, the basis for both CMOS and CCD sensors, is transparent above roughly 1100 nm, so imaging beyond that point must use a different detector material such as indium gallium arsenide, or InGaAs. However, current scientific-grade InGaAs cameras don’t offer the ultrafast imaging possible in the visible and near-infrared, where speeds can run in to the thousands of frames per second.
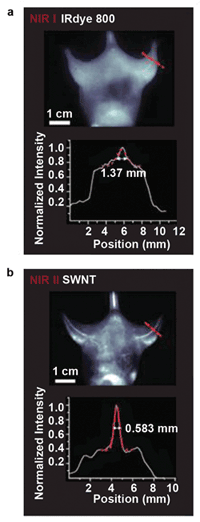
Imaging in the second IR window (below), running from 1000 to 1700 nm, improves resolution and detail clarity as compared to imaging in the near-IR below 1000 nm, but it does require using InGaAs instead of a silicon-based camera, which currently means a slower frame rate. Courtesy of Princeton Instruments and Hongjie Dai, Stanford University.
The issue is not the sensor material, according to Ravi Guntupalli, vice president of sales and marketing at Princeton Instruments. The Trenton, N.J.-based company produces a line of indium gallium arsenide cameras that run at over 100 fps.
The sensor is built in layers, with the top being an InGaAs absorption layer. This is bump-bonded onto a readout circuit made of silicon, yielding a device with an inherently parallel and potentially speedy readout.
Most of the demand for high-speed imaging comes from impact studies, which capture what happens when two objects slam together, according to Guntupalli. The parameters demanded by these applications differ from those needed by the life sciences.
“Biological applications require better sensitivity while preserving the frame rate. Achieving this combination is constrained by the current sensor-readout technology,” Guntupalli said.
He added that with speeds above 100 fps, Princeton Instruments’ existing InGaAs cameras can image events on a time frame of a few milliseconds with high sensitivity. Much higher speeds are possible below 900 nm, where silicon-based electron multiplying CCD (EMCCD) technology offers sensitivity and speed. If a full-resolution frame can be imaged 60 times a second, windowing that down to 16 × 16 pixels can up the speed to over 5000 fps, according to Guntupalli.
Whatever the wavelength or the sensor material used, it’s impossible to escape the laws of physics. Specifically, imaging at 1000-plus fps necessarily means that each frame is exposed for a millisecond or less. That limits the photons available for each frame.
“One of the challenges when running the camera faster is the significant decrease in the measured signal. Because of this, higher sensitivity becomes more critical in camera performance,” said Rachit Mohindra, product manager for Tucson, Ariz.-based Photometrics. The company is a sister firm to Princeton Instruments that makes ultraviolet, visible and near-IR scientific EMCCD and CCD microscopy cameras.
Certain areas of investigation, such as those involving processes in the heart or brain, could benefit from imaging in the 1000- to 2000-fps range. Photometrics cameras run at more than 500 fps but can be bumped up to three times that through the use of two techniques. One is windowing, which cuts down the number of pixels. Another is the creation of a superpixel, in which multiple pixels are binned and combined together. The advantage of this second approach is a higher signal-to-noise ratio and faster speeds; the downside is decreased resolution.
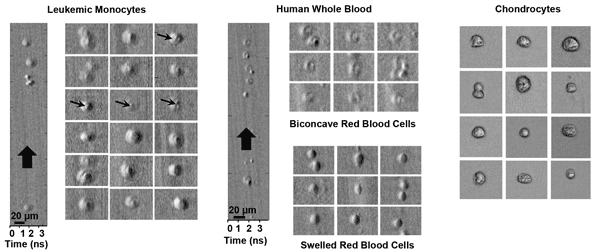
Leukemic monocytes, human whole blood and chondrocytes in an ultrafast microfluidic flow moving at roughly 10 m/s, imaged with ATOM, a second-generation time-stretch imaging technique. Courtesy of Kevin Tsia, University of Hong Kong.
Bio-based imaging tends to be photon starved, so anything that ups the signal can be highly advantageous. Given that, the capability of a detector is important. EMCCDs offer sensitivities of 95 percent in the visible and as much as 10 percent at 1000 nm, Mohindra indicated.
For an ultrafast imaging technique based on nontraditional methods, consider optical time-stretch imaging. Originally developed at the University of California, Los Angeles, it is now being further refined there and at the University of Hong Kong. In optical time-stretch imaging, a stream of femto- to picosecond broadband light pulses interrogates a sample. Image information is then extracted by time-stretching each individual output flash that results.
Heading the Hong Kong effort is Dr. Kevin Tsia, an assistant professor in the department of electrical and electronic engineering. Two key advantages of the technique are speeds in the millions of frames per second and enhanced sensitivity, Tsia said. The first arises because the frame rate is set by ultrafast image encoding into the optical pulse, which operates as fast as tens of megahertz. As for the second benefit, the encoded-image signal can be optically amplified, eliminating the typical trade-off of sensitivity for speed.

Sequentially timed image frames are encoded into ultrashort pulses in different spectral bands and digitally integrated into a movie. Consequently, sequentially timed all-optical mapping photography is capable of a movie capture rate of trillions of frames per second. Courtesy of Keisuke Goda, University of Tokyo.
In their latest efforts, the Hong Kong researchers have achieved high-contrast time-stretch images of label-free live cells, Tsia said. “Our results show that it can, for the first time, capture high-contrast and blur-free images of the ultrafast-moving stain-free mammalian living cells with subcellular resolution in microfluidic flow at an unprecedented speed of about 10 meters per second.”
That speed enables image-based single-cell screening and identification at a rate of 100,000 cells per second, compared with conventional techniques that are limited to about one-hundredth of that, he added. That faster screening capability could be of interest to those involved in single-cell analysis, drug discovery and rare cancer cell detection.
One of the fastest imaging methods currently possible is the sequentially timed all-optical mapping photography technique. Dr. Keisuke Goda, a University of Tokyo physical chemistry professor, was a co-author of an August Nature Photonics paper (doi: 10.1038/nphoton.2014.163) describing the technique.
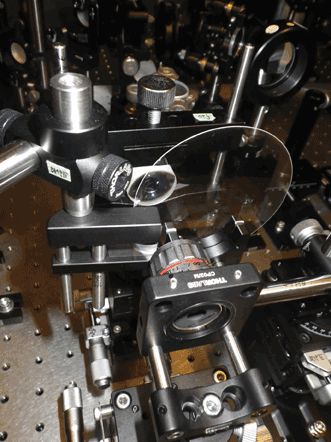

Pulses from this sequentially timed all-optical mapping photography camera illuminate a target with successive flashes. Image-encoded pulses are optically separate for simultaneous acquisition of multiple image frames. Courtesy of Keisuke Goda, University of Tokyo.
It splits ultrashort femtosecond laser pulses into daughter pulses in different spectral bands. These create consecutive flashes that are used for image acquisition – they are mapped into different areas of an image sensor. The result is an ability to capture 450 × 450-pixel images at an extremely fast rate.
“STAMP, or sequentially timed all-optical mapping photography, is an emerging enabling technology that enables burst-mode image acquisition at a frame rate of more than 1 trillion frames per second,” Goda said.
He listed a number of other non-CCD/CMOS sensor techniques that are being developed. Only with such emerging technologies can the limitations of conventional sensors be overcome, thereby producing the ability to screen large numbers of cells in an automated fashion, he added.
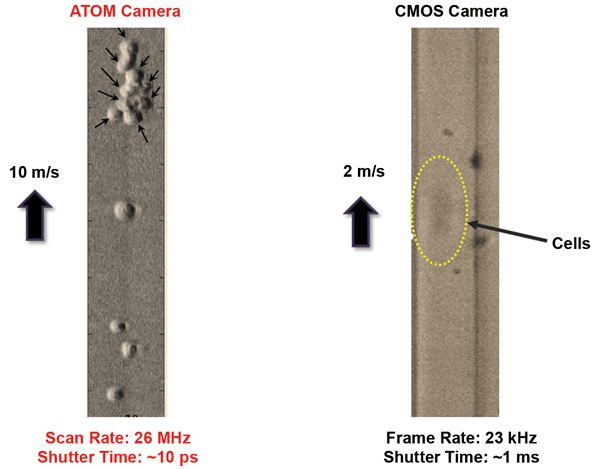

Imaging with ATOM, a second-generation time-stretch imaging technique, can resolve cells moving at a higher rate than is possible using a traditional state-of-the-art CMOS camera. Courtesy of Kevin Tsia, University of Hong Kong.
Finally, for any imaging technique, one common challenge is how to handle the flood of data generated. Collecting 4-MP images at 100 fps creates 800 MB of data per second. Even a short run can lead to tens of gigabytes of data, and such runs may be repeated many times a day. Thus, the storage system, network, image processing system and the rest of the back end must be built to handle this. Doing so without creating a bottleneck involves the use of high-capacity, expensive conventional high-speed hard disk drives or costly high-capacity solid-state drives, along with a hefty bandwidth network and computers with enough processing power.
“The industry is recognizing that there are more and more logistical challenges with the volume of data being generated,” said Photometrics’ Mohindra. “We are all working to address efficient and less costly ways for researchers to manage data.”