Dr. Marinella G. Sandros, University of North Carolina at Greensboro, and Dr. Fran Adar, Horiba Scientific
The first article in this two-part series looked at Raman spectroscopy applications in tumor surgery, endoscopy and bacterial identification. This second part examines applications in cell sorting, laser trapping, biology probing and more.
The field of medical diagnostics will have tremendous potential gains in the near future due to Raman spectroscopy and microscopy. Applications include surface-enhanced Raman spectroscopy (SERS), signal enhancement targeted to specific analytes, coupling atomic force microscopy (AFM) systems and enabling tip-enhanced Raman spectroscopy (TERS) for nanoscale resolution.
These technologies highlight great research developments that we anticipate will be transitioning sooner rather than later.
Laser trapping
Laser trapping refers to a well-focused laser beam’s ability to trap a microscopic dielectric particle by balancing the optical gradient forces in the focused beam and the transfer of photon momentum. Such trapping enables the stable positioning of particles for Raman analysis. Additionally, if they are trapped in an aqueous stream, they are presented to the measurement in their native state and cooled by the flowing liquid.
The earliest articles describing laser trapping in combination with Raman measurements of which we are aware were published in 1996,1 1997,2 20013 and 2003.4 Using this technology, Xie et al5 reported the ability to monitor real-time changes during heat denaturation. Several years later, they reported the ability to identify species by measuring single laser-trapped bacteria suspended above the sampling substrate.5
An alternative method for trapping particles – trapping in the evanescent field outside waveguides6 – was developed using near-field photonics. The argument for using this method is that it produces the higher optical forces necessary for capturing particles on the nanoscale. This technique has been commercialized for trapping submicrometer particles flowing above the waveguide. By using a 785-nm laser in the waveguide, it is possible to simultaneously capture the Raman signal simply by mounting the chip on a standard Raman microscope.7
Another flow-through lab-on-a-chip was developed for studying tumor cells flowing and/or trapped in a reservoir;8 a dual-beam optical trap was constructed with fiber lasers. This technology was further developed with the addition of fiber Bragg gratings for collecting Raman light; by eliminating the need for a microscope and high-numerical-aperture objectives, the systems can be simplified and made less costly. All of these devices that can control particle flow before and after Raman measurement have the potential to subsequently provide cell sorting.
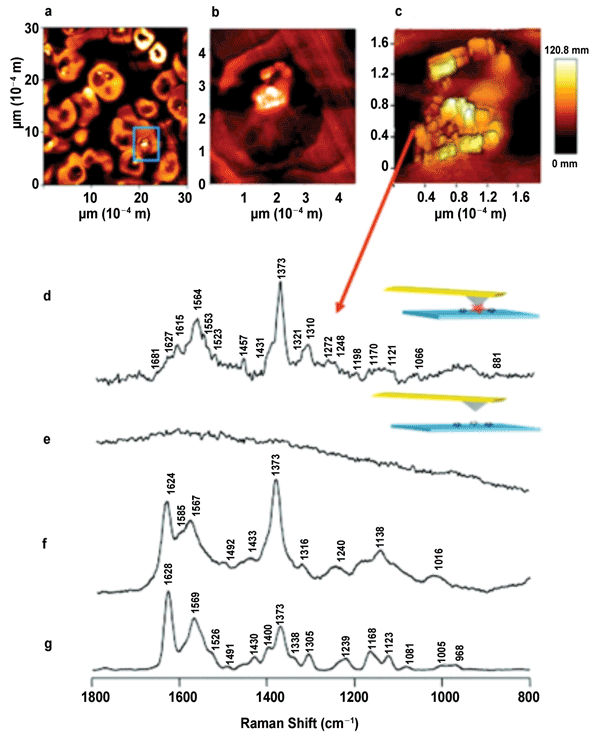
Figure 1. (a-c) AFM images the localization of hemozoin crystals inside
sectioned malaria-infected red blood cells. (Below) Tip-enhanced Raman
spectroscopy spectrum representing (d) the edge of a hemozoin crystal,
(e) the tip lifted up away from the sample several microns, (f) SERS
analysis of β-hematin (synthetic analogue of hemozoin), (g) resonance
Raman spectrum of β-hematin. Adapted with permission from B.R. Wood et
al (2011). Tip-enhanced Raman scattering (TERS) from hemozoin crystals
within a sectioned erythrocyte. Nano Lett, Vol. 11, pp. 1868-1873. ©
2011 American Chemical Society.
Cell sorting
Raman-activated cell sorting has been demonstrated on a microfluidic chip with optical tweezers in which cells were captured, analyzed and sorted.9,10 The advantage of this method is that the cell is left intact and available for further processing.
Raman methods to decrease the acquisition time (and thus the decision making time), including SERS, coherent anti-Stokes Raman scattering and stimulated Raman scattering, have been reviewed.11 In fact, the same authors showed the possibility of recording spectra in as little time as one-tenth of a second on a modified instrument.12 In that article, they studied bacteria isolated from the oral cavity. They were successful in classifying five types of bacteria and in differentiating cells that had incorporated 13C and 15N from isotopically labeled nutrients. Based on these identification abilities, they implemented Raman-activated cell ejection assisted by laser-induced forward transfer. This technology can now sort single cells in complex samples such as biofilms, soils, sludge and tissue. In a subsequent publication,13 this group showed that culturing and genomic sequencing of isolated cells enable new organisms to be identified. And the isotope probing enables studying genetic function and physiology of unculturable organisms in the ecosystem.
SERS and TERS probing biology
Disease detection at the early stage is often challenging, as biomarker expression is very low and spontaneous Raman microscopy does not provide the sensitivity needed to highlight its presence in biological systems. To amplify the Raman vibrational signals, scientists turned to SERS. This is achieved either by detecting the biomarker directly, or by detecting a tag for a specific biomarker. Direct SERS analysis occurs when a biomarker is in proximity (<10 nm) to a noble-metal (i.e., gold or silver) substrate. The indirect mode uses a reporter molecule as a “tag” that will highlight the presence of a specific biomarker. The SERS probe is constructed by depositing a self-assembled monolayer of reporter molecules around the metal nanoparticle and then coating it with a polymer to stabilize and facilitate attachment of antibodies or aptamers that enhance target selectivity.14 SERS enhancement can increase signal by factors of 106 to 1014,15 through the excitation of localized surface plasmon resonances16,17 arising from the metal nanoparticle surface.
To date, an explosion of publications has highlighted how the coupling of SERS18 with Raman spectroscopy and microscopy can advance measurement of biological activity. By using a SERS probe, Bowey et al19 showed that one can pinpoint and quantify the expression of a biomarker at the single-cell level; they also showed that different types of tags can be used to detect multiple biomarkers simultaneously.20 Furthermore, single-biomolecule sensing was achieved by using a unique construct of gold nanospheres on gold thin film.21
A comparative study by McQueenie et al22 examined SERS performance in detecting an inflammatory biomarker with conventional fluorescence imaging in vitro, ex vivo and in vivo. Unanimously, the assessment showed that SERS provides better signal-to-noise ratio, narrower spectral features, greater penetration depth and improved sensitivity compared with current systems. An interesting study by Premasiri and colleagues23
reported that SERS analysis can also reveal the presence of hypoxanthine (a biomarker for purine metabolism disorders such as gout24), which was not present in the normal Raman spectrum of human whole blood. Furthermore, they indicated that Raman signal of hypoxanthine changed over time. This work exemplifies the potential of SERS to understand the underlying biochemical mechanism responsible for this type of fluctuation. In addition, Premasiri’s group currently is working on developing a robust database for diagnosing urinary tract infection pathogens using SERS.
Despite all the interesting publications with SERS, some challenges still need to be addressed before it transitions to a clinical setting. One major issue is irreproducibility, which occurs because the method for the fabrication of SERS substrate or particles does not always generate the same signal-enhancing “hot spots.” It is foreseen that, in the future, smart-surface nanoengineering will play a big role in improving SERS applications.
Before closing, we should describe TERS,25 which integrates AFM with Raman microscopy, offering the sensitivity of SERS along with the ultrahigh spatial resolution provided by the AFM tip. This technique has been shown to identify molecular information at the nanoscale for the detection of different virus strains,26 peptide nanotapes,27 DNA sequencing28 and formation of hemozoin crystal within individual red blood cells infected with the malaria parasite,29 to name a few. As evidence of this technology’s value, Figure 1 shows Raman images of hemozoin crystals (at 20-nm spatial resolution) that were found within the digestive vacuole of a sectioned red blood cell. The next step with this platform is to screen drugs that could bind to hemozoin in the hopes of improving treatment for malaria.
This article presents a few examples highlighting the rewards afforded by Raman technology in combination with ultrasensitive nanomaterials and sophisticated tools like optical tweezers and AFM. These applications are still new and require some fine-tuning; however, we foresee that they will lead to new applied life science discoveries that potentially could impact unmet medical solutions.
Meet the authors
Dr. Marinella G. Sandros is an assistant professor in the Department of Nanoscience at the University of North Carolina at Greensboro; email: [email protected]. Dr. Fran Adar is the principal Raman applications scientist at Horiba Scientific in Edison, N.J.; email: [email protected].
References
1. C. Esen et al (1996). Raman investigation of photopolymerization reactions of single optically levitated microparticles. Appl Spectrosc, Vol. 50, pp. 823-828.
2. M. Trunk et al (1997). Microchemistry: time dependence of an acid-base reaction in a single optically levitated microdroplet. Chem Phys Lett, Vol. 264, pp. 233-237.
3. K. Ajito and K. Torimitsu (2001). Near-infrared Raman spectroscopy of single particles. TrAC, Trends Anal Chem, Vol. 20, pp. 255-262.
4. C. Xie et al (2003). Study of dynamical process of heat denaturation in optically trapped single microorganisms by near-infrared Raman spectroscopy. J Appl Phys, Vol. 94, pp. 6138-6142.
5. C. Xie et al (2005). Identification of single bacterial cells in aqueous solution using confocal laser tweezers Raman spectroscopy. Anal Chem, Vol. 77, pp. 4390-4397.
6. D. Erickson et al (2011). Nanomanipulation using near field photonics. Lab Chip, Vol. 11, pp. 995-1009.
7. http://www.opfluid.com/#!nanoparticle-identification/c1ygk.
8. S. Dochow et al (2013). Quartz microfluidic chip for tumour cell identification by Raman spectroscopy in combination with optical traps. Anal Bioanal Chem, Vol. 405, pp. 2743-2746.
9. A. Lau et al (2008). An integrated optofluidic platform for Raman-activated cell sorting. Lab Chip, Vol. 8, pp. 1116-1120.
10. W.E. Huang et al (2009). Raman tweezers sorting of single microbial cells. Environ Microbiol Rep, Vol. 1, pp. 44-49.
11. M. Li et al (2012). Single-cell Raman spectroscopy for cell sorting and imaging. Curr Opin Biotechnol, Vol. 23, pp. 56-63.
12. Y. Wang et al (2013). Raman-activated cell ejection for isolation of single cells. Anal Chem, Vol. 85, pp. 10697-10701.
13. M. Li (2014). Single-cell Raman sorting. Environ Microbiol. 2nd ed. I.T. Paulsen and A.J. Holmes, eds. Humana Press, pp. 147-153.
14. S. Schlücker (2009). SERS microscopy: nanoparticle probes and biomedical applications. ChemPhysChem, Vol. 10, pp. 1344-1354.
15. S. Nie and S.R. Emory (1997). Probing single molecules and single nanoparticles by surface-enhanced Raman scattering. Science, Vol. 275, pp. 1102-1106.
16. E.C. Le Ru and P.G. Etchegoin (2009). Principles of surface-enhanced Raman spectroscopy and related plasmonic effects. Elsevier, Amsterdam.
17. E.C. Le Ru and P.G. Etchegoin (2011). Basic electromagnetic theory of SERS, in surface-enhanced Raman spectroscopy: Analytical, biophysical and life science applications. Wiley-VCH, Weinheim, Germany.
18. S. Efrima and L. Zeiri (2009). Understanding SERS of bacteria. J Raman Spectrosc, Vol. 40, pp. 277-288.
19. K. Bowey et al (2014). Microwave-assisted synthesis of surface-enhanced Raman scattering nanoprobes for cellular sensing. Colloid Surface B, Vol. 122, pp. 617-622.
20. I. Kim et al (2012). SERS-based multiple biomarker detection using a gold-patterned microarray chip. J Mol Struct, Vol. 1023, pp. 197-203.
21. L. Li et al (2013). Single molecule SERS and detection of biomolecules with a single gold nanoparticle on a mirror junction. Analyst, Vol. 138, pp. 4574-4578.
22. R. McQueenie et al (2012). Detection of inflammation in vivo by surface-enhanced Raman scattering provides higher sensitivity than conventional fluorescence imaging. Anal Chem, Vol. 84, pp. 5968-5975.
23. W.R. Premasiri et al (2012). Surface-enhanced Raman scattering of whole human blood, blood plasma and red blood cells: Cellular processes and bioanalytical sensing. J Phys Chem B, Vol. 116, pp. 9376-9386.
24. T. Yamamoto (1995). Determination of plasma purine nucleoside phosphorylase activity by high-performance liquid chromatography. Anal Biochem, Vol. 227, pp. 135-139.
25. R.M. Stöckle et al (2000). Nanoscale chemical analysis by tip-enhanced Raman spectroscopy. Chem Phys Lett, Vol. 318, pp. 131-136.
26. P. Hermann et al (2011). Evaluation of tip-enhanced Raman spectroscopy for characterizing different virus strains. Analyst, Vol. 136, pp. 1148-1152.
27. M. Paulite et al (2013). Full spectroscopic tip-enhanced Raman imaging of single nanotapes formed from β-amyloid (1-40) peptide fragments. ACS Nano, Vol. 7, pp. 911-920.
28. R. Treffer et al (2011). Distinction of nucleobases – a tip-enhanced Raman approach. Beilstein J Nanotechnol, Vol. 2, pp. 628-637.
29. B.R. Wood et al (2011). Tip-enhanced Raman scattering (TERS) from hemozoin crystals within a sectioned erythrocyte. Nano Lett, Vol. 11, pp. 1868-1873.