Emine Korkmaz, FEI
In recent years, there have been considerable advancements in EM-based methods for 3-D reconstructions of large tissue volumes. In this article, we describe a new approach that combines in situ serial sectioning with a unique, multienergy deconvolution technique to generate 3-D reconstructions with isotropic resolution down to a few nanometers.
Unraveling complex 3-D architecture of cells and tissues in their natural context is crucial for understanding relationships between structure and function in biological systems. Since its invention, light microscopy (LM) has been an indispensable tool for these biological investigations and it has many advantages within this context. For instance, it requires minimal sample preparation and it can be used to look at living samples. Also, many biological samples are nearly transparent to visible light, permitting relatively easy 3-D analysis. However, LM resolution fundamentally is limited by the wavelength of visible light, allowing it to resolve structural details no smaller than several hundred nanometers. In its various methods of use, electron microscopy (EM) can resolve details down to the atomic scale, although practical considerations generally limit its resolution to a few nanometers in most biological applications. Still, this is sufficient to resolve many of the structures of interest in current biological investigations. Bulk biological materials, however, are not transparent to electrons, making 3-D imaging and analysis significantly more challenging.
Imaging the same sample with both LM and EM techniques provides vital information from different modalities. Correlative light and electron microscopy (CLEM) is a powerful tool used to combine the study of dynamic processes or rare events by using LM to screen and identify specific regions of interest at a large scale of cells or tissues. Linking this functional information to high-resolution, ultrastructural information by way of 3-D EM techniques, such as serial block-face scanning electron microscopy (SBF-SEM), presents new avenues to expand ultrastructural description to functional context with the flexibility of choosing the volume size for a more comprehensive understanding of complex biological events. This approach demands a compatible sample preparation technique that facilitates the application of both LM and EM on a bulk biological sample.
EM-based 3-D reconstruction techniques
A transmission electron microscope (TEM) forms an image from electrons that have passed through the sample. Because electrons interact strongly with the sample and electron transmission is required for image formation, it must be very thin – typically less than 100 nm for an 80- to 120-keV electron beam. Images collected from a series of thin sections can be computationally combined to generate a 3-D model of the sectioned volume (Figure 1). TEMs are capable of very high lateral (X-Y) resolution. However, Z-resolution in this type of reconstruction is determined by the thickness of the sections, which is practically limited to 40 to 50 nm. It is possible to improve the Z-resolution of such an approach by using electron tomography to image each section. Electron tomography reconstructs a 3-D structure from a series of images acquired from different perspectives. While it is theoretically capable of nearly isotropic resolution, the additional imaging and reconstruction of each section adds considerable time and difficulty to the overall reconstruction process.
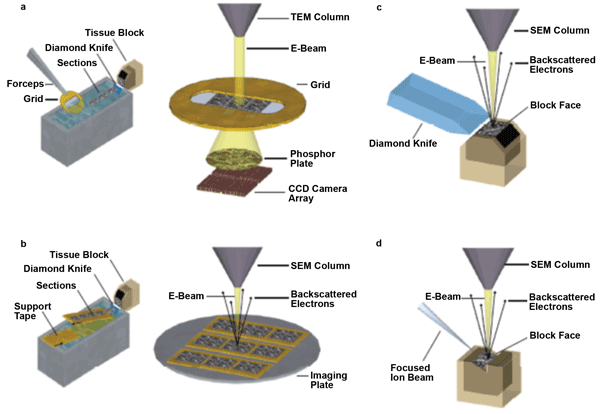
Figure 1. Simplified schematics (not to scale) of the volume EM techniques described. (a) Serial section transmission electron microscopy (ssTEM). Sections are cut by hand with an ultramicrotome, floated onto a water boat and then picked up onto grids. In ssTEM, electrons transmitted through the sample are focused with electron optics (not pictured) to form an image on a phosphor plate. The image is recorded digitally with, in this case, a CCD TEM camera array (TEMCA). (b) Automated tape-collecting ultramicrotome scanning electron microscopy (ATUM-SEM). Sections are cut automatically on an ultramicrotome and collected from the water bath using a custom-designed, tape-collection conveyor belt. Because the support tape is electron opaque, ATUM sections are mounted on an imaging plate and imaged in a SEM. Images are formed by collecting backscattered electrons with an electron detector mounted above the sample (not shown). (c) Serial block-face scanning electron microscopy (SBF-SEM). Automated sectioning with a diamond knife and imaging are performed within the vacuum chamber of a SEM using a custom-designed microtome and specimen stage. (d) Focused ion beam milling scanning electron microscopy (FIB-SEM). Automated milling with a focused ion beam and imaging are performed within a dual-beam SEM.2
SEM scans a finely focused electron beam in a raster pattern over the surface of a bulk specimen and creates a virtual image by correlating changes in various signals with the position of the beam in the raster. The most useful signal for 3-D imaging is the backscattered electron (BSE) signal, which reflects the average atomic number of the region illuminated by the beam. BSEs are beam electrons that are scattered back out of the sample surface by interactions with nuclei of sample atoms. They originate from a volume of interaction located immediately below the beam – the depth of that volume is determined by the beam’s energy. Biological electron microscopists have developed a number of staining techniques to enhance BSE contrast by selectively staining certain structures with heavy elements.
SEM-based methods may involve the imaging of serial sections, which are automatically collected on an electron-opaque tape – similar to the TEM-based serial sectioning method described above. Alternatively, these methods may be based on images of the surface of a plastic-embedded block specimen, known as serial block-face (SBF) imaging.1,2 SBF involves imaging of the block face followed by the removal of a thin layer of material. This sequence is repeated many times to generate a stack of images from which a 3-D structure can be reconstructed. As with the other serial sectioning techniques, the Z-resolution primarily is determined by the thickness of the sections. This can be thinner because the structural integrity of the removed layer is not required to be preserved for subsequent imaging. Still, imaging is limited to 20 to 30 nm due to mechanical sectioning techniques. The thickness of the section can be reduced significantly by using a focused ion beam (FIB) for the removal, yielding a nearly isotropic resolution. FIB sectioning, however, is much slower than mechanical sectioning.
Recently, in-chamber ultramicrotomes have become available that can automatically remove thin layers from the block face for SBF reconstructions. These simplify the SBF process but they still are limited to removing layers with a practical minimum thickness of about 25 nm. While an in-chamber microtome can accelerate the sectioning process, it alone does not address other challenges of SBF, such as the management of charging artifacts and contrast changes associated with variations in staining. It also does not address the computational aspects of stitching (used to image larger areas), reconstruction, visualization and analysis.
Combining physical and optical sectioning
FEI’s recently introduced solution, Teneo VS, overcomes the Z-resolution limit by combining mechanical sectioning with an optical sectioning technique known as multienergy deconvolution (MED), as is indicated in Figure 2. The size and shape of the volume of interaction (VOI) below the beam spot, from which BSE originates, varies with the energy of the electron beam and the composition of the sample. Higher-energy electrons penetrate more deeply into the sample. In heavy materials, the increased penetration is accompanied by a broadening of the VOI, which results in a loss of lateral resolution. However, in biological samples, which are predominately composed of lighter elements, this broadening is much less pronounced and the lateral spread of the interaction volume is limited.

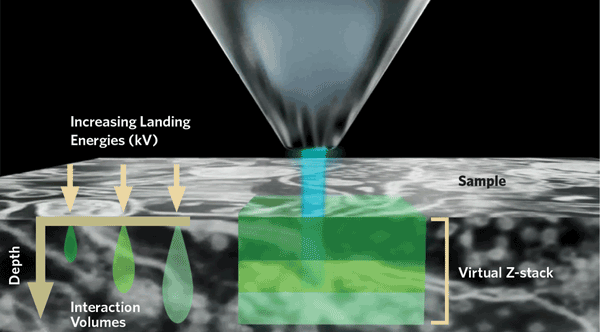
Figure 2. Multienergy deconvolution acquires several images at different accelerating voltages. Each image contains information about a layer of different thickness. A proprietary software algorithm deconvolutes the data to provide a subset of optical sections for each physical section. The final reconstructed model exhibits isotropic resolution down to 10 nm.
The SBF-MED process begins with physical sectioning by an in-chamber microtome using a diamond knife. The microtome itself mounts to the SEM stage and is easily removable to permit use of the SEM for non-SBF applications. The microtome removes layers that range from 40 to 50 nm in thickness. After each physical section, the MED process acquires multiple images of the block face at varying beam energies. It then applies a proprietary deconvolution procedure to the images to compute optical sections at various depths within the physical section, subsequently forming a 3-D subset. Combining the subsets for each of the physical sections results in an overall
reconstruction that exhibits isotropic resolution. Essentially, this results in equivalent resolution in all directions.
SEM image acquisition is a serial process, and requirements for resolution and signal-to-noise ratio (SNR) fundamentally determine the time required to acquire an image. Consequently, the strength of the signal and the efficiency are important considerations. The strength of the signal is determined primarily by the amount of beam current the sample can tolerate before damage occurs – biological samples, even when embedded, can be especially sensitive to beam damage. Therefore, the detection system needs to be optimized to collect low-loss BSE signal with high efficiency, which is best done by using an in-lens detection geometry that generates images with strong contrast and high SNR.
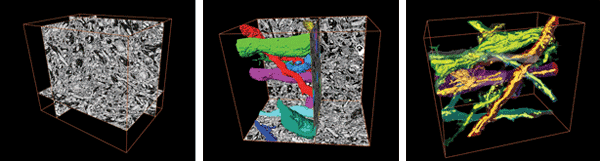

Figure 3. Volume reconstruction of a mouse brain acquired with a combination of SBF-SEM and MED-SEM at HiVac-mode. The block face was imaged in BSE mode at varying accelerating voltages, ranging from 1.2- to 3.1-kV isotropic resolution of 10 × 10 × 10-nm pixels (X, Y, Z), 1040 slices (physical + optical slices) at 50 nm (physical slices), volume 15.00 µm × 12.9 µm × 10.4 µm (1040 slices) or 18.7 µm × 15.5 µm × 6.8 µm (680 slices). Courtesy of P. Laserstein and P. Bastians, Helmstaedter Lab, MPI Brain Research, Germany.
Charging artifacts occur in SEM images when charges accumulate on nonconductive samples. Plastic-embedded biological samples can be very difficult to image. In routine imaging applications, conductive coatings may be applied to nonconductive samples to provide a path to ground the electrical current and thereby prevent charge accumulation. Coatings cannot be used in SBF where a continuously sectioned block-face surface is exposed to the beam each time a layer of material is removed by the in-chamber microtome. Low-vacuum (LV) operation provides a means to eliminate charging that is compatible with SBF. In LV mode, a gas – in this case, water vapor – is injected into the sample chamber and maintained at a pressure low enough to permit the beam to reach the sample without excessive scattering. Some molecules of the gas are ionized by beam electrons or by electrons originating from the sample. These ion gas molecules are then available to neutralize any charge that accumulates on the sample. LV operation is essential to ensure high-quality imaging of difficult samples.
As data sets become larger and data acquisition times increase, automation becomes necessary to ensure productivity. In the system described here, the entire SBF process is automated and controlled by an integrated user interface and software-based automation. FEI’s MAPS software automates processes ranging from low-level setup and column alignment up through walk-away image series acquisition. The software also provides a means to acquire data from block faces larger than a single field of view by stitching together multiple images. Additionally, it can overlay images from other techniques, such as fluorescent light microscopy, to permit fast and reliable navigation to a feature of interest identified by specialized labeling techniques.
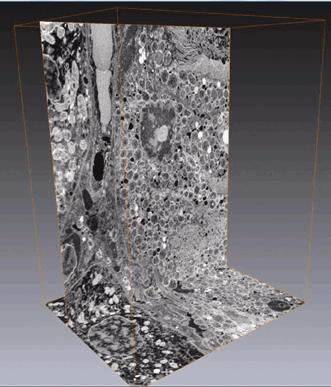

Figure 4. Volume reconstruction of a mouse kidney acquired with SBF-SEM at LoVac mode. The block face was imaged in BSE mode at 3-kV accelerating voltage, 0.2-nA beam current, 5 × 1-μs dwell time, 8 × 8-nm pixels (X,Y), 554 slices at 50 nm, volume 28.7 × 25.9 × 27.7 μm. Courtesy of FEI.
Reconstruction, visualization and analysis are provided by our Amira software platform, which is an extendable software system for 3-D and 4-D analyses that can be used in many applications. The platform also includes tools for image segmentation and geometry reconstruction. This allows the user to mark, or segment, structures and regions of interest in 3-D image volumes using automatic, semiautomatic and manual tools. The segmentation then can be used for a variety of subsequent tasks including volumetric analysis, shape analysis and analyzing computer-generated 3-D models. Other capabilities include multiplanar and volume visualization, image registration, filament tracing, and cell separation and analysis.
We have presented a novel SBF solution that combines mechanical sectioning with multienergy deconvolution to provide isotropic resolution not achievable with mechanical sectioning alone. Optimized detectors and low-vacuum operation ensure high-quality imaging on all biological samples. Software provides extensive automation of processes, ranging from low-level setup and alignment to walk-away acquisition of the complete image series. We also have described software solutions for large-area and large-volume analysis, reconstruction and visualization. The full integration of all SBF imaging capabilities ensures ease of use and flexibility for various samples and experiments.
Meet the author
Emine Korkmaz is product marketing manager for life sciences scanning electron microscopy and large-volume EM imaging at FEI in Eindhoven, the Netherlands. Email: [email protected].
References
1. W. Denk and H. Horstman (2004). Serial block-face imaging scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLOS Biol, Vol. 2.
2. K. Briggman and D. Bock (2012). Volume electron microscopy for neuronal circuit reconstruction. Curr Opin Neurobiol, Vol. 22, pp. 154-161.