Three-dimensional holographic microscopy enables quantitative and noninvasive investigation of cells’ intrinsic properties — without labeling agents. The technique could prove invaluable in cancer diagnosis, treating infection and hematology.
JODI POUNDS, TOMOCUBE INC
Cellular analysis plays a crucial role in a wide variety of research and diagnostic activities in the life sciences. However, the information available to researchers and clinicians is limited by current microscopy devices and techniques. An innovative new tool — a microscope that displays 3D holograms — can overcome many of these limitations and open new vistas for researchers and clinicians to understand, diagnose and treat disease.
The word hologram — from the Greek “holos,” meaning “whole,” and “graphe,” meaning “to write”— describes the completeness of cellular information provided by holographic microscopy. This is the first technique that allows the observation of living cells in three dimensions so users can visually travel around the exterior and through the interior of a cell.
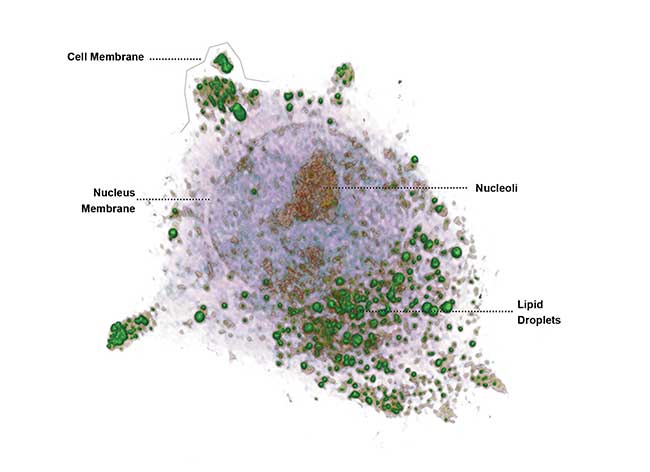

Hepatocyte detail captured by Tomocube Inc.’s HT-1 holographic microscope. Courtesy of Tomocube Inc.
This technique utilizes optical diffraction tomography (ODT), which enables users to quantitatively and noninvasively investigate the intrinsic properties of cells. ODT reconstructs the 3D distribution of the refractive index (RI) and by doing so, provides structural and chemical information of the cell including dry mass, morphology and dynamics of the cellular membrane. The images are incredibly precise — at a resolution of 110 nm, approximately 1,000 times smaller than the diameter of a human hair. The HT-1 3D holographic microscope, developed by YongKeun (Paul) Park, associate professor of physics at the Korea Advanced Institute of Science and Technology (KAIST), and commercialized by Tomocube Inc., both of Daejeon, South Korea, is one of only two microscopes of its kind available in the world today.
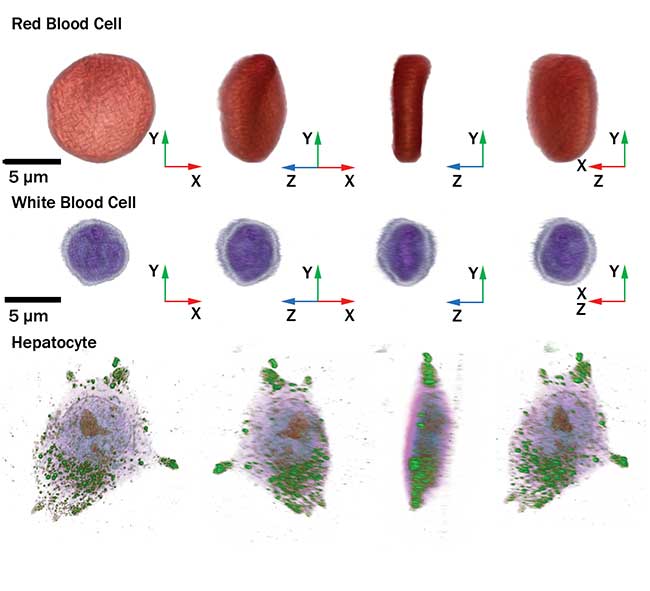
Cellular images captured by the HT-1 holographic microscope. Courtesy of Tomocube Inc.
State-of-the-art microscopy
Current alternative cell-imaging tools include fluorescent confocal and phase contrast microscopy. Both techniques have advantages for researchers in certain applications, but neither technique can provide high-resolution, 3D images of live cells.
Fluorescence confocal microscopy techniques require the use of labeling agents in order to render high-contrast molecular information. These agents can cause photobleaching, phototoxicity and interference with normal molecular activities. Cells that need to be analyzed and then reinserted into the body, such as stem or immune cells, pose an especially difficult challenge for confocal microscopy. The labeling process is also time-intensive and requires experts to spend hours preparing a sample for analysis.

Tomocube’s 3D holographic microscope, the HT-1 series. Courtesy of Tomocube Inc.
Phase-contrast microscopes address some limitations of fluorescent microscopy by eliminating the need for fluorescent stains that kill cells. These microscopes provide more information than a simpler bright-field microscope by measuring the variations in light passing through a transparent specimen. The changes in amplitude and phase of light waves depend on the medium through which the light is passing. By separating the background light from the scattered light from the specimen, the phase-contrast microscope can render detailed 2D images of living cells. This advancement earned its inventor, Frits Zernike, the Nobel Prize in physics in 1953.
However, the phase-contrast microscope does not provide quantitative information about cells, such as height, volume, thickness and mass. Attempts in the field of microscopy to provide quantitative information about cells have resulted in the scanning electron microscope and the atomic force microscope. Both alternatives successfully provide quantitative information about cells, but cannot image living cells.
Holotomography was developed in order to combine the advantages of the best currently available microscopy techniques: quantitative imaging and live-cell imaging. The principles behind the technique are relatively straightforward, but the results are groundbreaking.
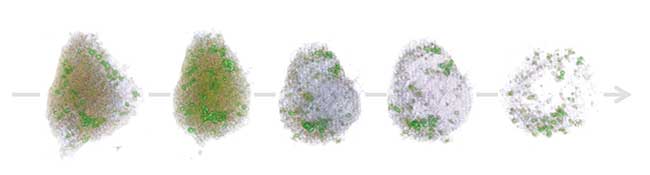
Hepatocyte in necrosis captured in 4D by the HT-1 holographic microscope. Courtesy of Tomocube Inc.
How holotomography works
All objects can be measured by a refractive index distribution, which is an intrinsic optical parameter determined by the speed at which light passes through the specific material. The light passing through the cell and its component parts is slower than the light passing through the medium surrounding it. By employing interferometrics, one can retrieve both phase and amplitude to render a topogram, or a 2D phase delay map.
In order to measure a 3D tomogram, a technique analogous to an x-ray computed tomography (CT) system is employed. An x-ray CT measures 3D distributions of x-ray absorbtion parameters of the human body, and displays images of human organs and tissues. X-ray CT utilizes a rotating energy source to capture multiple 2D images of the human body, from which 3D tomograms of the absorption parameters are retrievable.
The same principle is utilized in 3D holotomography. By scanning the illumination angle of the beam, and measuring multiple 2D holograms, enough data is generated to then reconstruct a 3D image using an appropriate algorithm. The process of digital reconstruction is a difficult one and current work builds on the advancements made by physicist Emil Wolf in 1969.
Generally, there are two options for algorithmic reconstruction of a 3D cell image from multiple 2D RI topograms: a projection algorithm and a diffraction algorithm. The projection algorithm is analogous to x-ray computerized tomography and is calculated via the filtered back-projection method1. The method works best for samples that are nearly transparent. A diffraction algorithm accounts for the diffraction of light induced by the sample and displays high image quality with less distortion.
The instrumentation setup is partially dependent on the application and setting, as well as user preferences, and a variety of schematics have been deployed. However, the main attributes for holotomography are consistent: Using laser interferometrics, one laser beam is refracted from the sample, which yields amplitude and phase information about the sample. By changing the illumination angle, multiple 2D RI measurements are then stitched together using a reconstruction algorithm to display a 3D image on the user’s computer screen. RI (n) values describe how light propagates through a medium and are defined as n = c/v where c is the speed of light in a vacuum and v is the phase velocity of light in the medium.
Because the RI values are a quantitative measurement, this technique has the significant advantage of providing quantitative data about the sample, including morphological and chemical information. Unlike other microscopy techniques like fluorescent confocal microscopy, this quantitative measurement of RI is not dependent on sample preparation or experimental setup. So, the instrumentation provides reproducible data for researchers and clinicians.
Park’s team has advanced the art by utilizing a specialized technique for capturing optical diffraction. In order to retrieve holotomographic information, the illumination beam must be passed over the sample in 360 degrees. Previously, approaches have included using a galvanometric mirror to interfere with the illumination beam, sending it through the sample at a variety of angles, or utilizing a mechanical device to rotate the sample itself while the illumination beam remains static. Neither technique is ideal for reliable, commercially available devices. Galvanometric mirrors are very costly and require regular calibration and alignment, making them impractical for regular use. And rotating a biological sample is also problematic as the soft and elastic biological material can be altered by the force of the mechanical rotation.
To address those limitations, Park’s team deployed a dynamic micromirror device (DMD), effectively eliminating moving parts from their microscope. A DMD is made up of many individually controlled and switchable micromirrors that can each be individually controlled. A DMD can be used to control the illumination angle in diffraction tomography to provide electronic control for better stability and improved optical resolution. The principle behind this is to utilize grating to diffract the laser. The angle of diffraction will be determined by the period of the grating. So, by controlling the period of the grating pattern, the diffraction angle of light can be controlled.
Hematological applications
Over the past several years Park and other researchers around the world have deployed holotomographic techniques in the life sciences, with publications describing applications in single-cell analysis, cytology and cytopathology, drug and toxicity reactions, and others. Another field in which holotomography could help advance disease diagnosis and treatment is in hematology. Information about red blood cells (RBCs) plays a critical role in hematology. In order to understand the pathophysiology of a number of diseases, such as malaria and sickle cell anemia, and to potentially diagnose these diseases earlier, it is crucial to be able to measure the morphological, chemical and mechanical properties of individual RBCs. However, current techniques, including optical microscopy and automated blood cell counters, can only provide limited information about the cells and lack the detail needed to properly observe the pathophysiologyof RBCs.
To better understand, diagnose and treat many blood-related diseases, a single-cell technique may be the answer. No current approaches to single-cell profiling provide complete characterization of individual RBCs. Recently, holotomography was used to demonstrate simultaneous characterization of the morphological, chemical and mechanical parameters of individual RBCs2. The research team, including Park, measured the 3D RI and 2D dynamic fluctuations of the cell membranes of individual red blood cells at the same time. This yielded the volume, surface area, sphericity, hemoglobin content and hemoglobin concentration. These factors can be important indicators for diagnosis of blood diseases and infections, and because they are measured quantitatively they can be further analyzed for better understanding of the causes and development of disease. The method can be deployed in diseases ranging from sickle-cell anemia to malaria.
Hematology is only one of a virtually limitless array of applications for this cell-imaging tool. The tool can also be used to provide instant quantitative cellular information to aid in cancer diagnosis, identify microbial infections, determine whether a particular treatment intervention has been successful, and address other research and medical challenges.
Meet the author
Jodi Pounds advises early stage technology companies in the areas of intellectual property and business development, helping clients commercialize their best ideas; email: [email protected].
References
1. K.R. Lee and K. Kim, et al. (2013). Quantitative phase imaging techniques for the study of cell pathophysiology: From principles to applications. Sensors, Vol. 13, pp. 4190-4191.
2. Y. Kim and H. Shim, et al. (2014). Profiling individual human red blood cells using common-path diffraction optical tomography. Sci Rep, Vol. 4, pp. 6659.