
3D Mapping of Neural Circuits In Vivo Opens the Window on Neurological Disease
Modifications with SLMs to existing two-photon microscopes can provide noninvasive probes deep within the cortex.
ANNA LINNENBERGER, MEADOWLARK OPTICS INC.
Despite extensive research, brain function and neurological diseases are poorly understood. Complexities arise from the quantity of neurons in the brain and from the densely interconnected networks of intermixed cell types. Tools neuroscientists have traditionally relied upon include the patch clamp, which probes electrical activity of a single neuron, and fMRI, which images activity in volumes containing millions of neurons.
These approaches target two vastly different scales. However, it is possible that the brain functions through firing patterns in neural circuits and that neurological disease is the result of alterations to the physical structure of circuits or circuit dynamics. These circuits exist at an intermediate scale that neither patch clamp nor fMRI can readily address. In order to give neuroscientists a range of tools to study brain function, there is a need for methods that noninvasively probe the underlying microcircuitry in the brain with single-cell resolution.
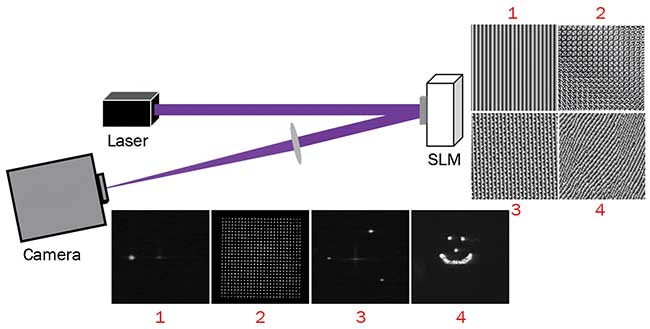
Figure 1. By manipulating the wavefront of a single incident beam, the spatial light modulator (SLM) can be used to superimpose lens and grating functions with weighting functions to redirect light to arbitrary locations to simultaneously create hundreds of focal points within a 3D volume. Courtesy of Meadowlark Optics.
Over the last decade, calcium imaging and photoactivation have emerged as solutions to this problem, providing all-optical means to monitor and manipulate circuit activity1. Calcium imaging uses calcium indicators that bind with calcium to alter the fluorescence characteristics of neurons. When a neuron fires, there is an uptake of calcium into the cell body. If the firing neuron is illuminated with an excitation source during the firing event, then the fluorescence emission increases, generating an optical response that corresponds to electrical activity.
Complementary to calcium imaging is photoactivation, which can use photosensitive proteins (optogenetics) or opto-chemical (caged) compounds to manipulate firing patterns either by causing neurons to fire or by silencing neurons. This combination of calcium imaging and photoactivation offers a means for neuroscientists to record the spatiotemporal dynamics of activity and map physical structure of circuits with single-cell resolution. However, without advanced microscopes for neuroscience, the benefits of calcium imaging and photoactivation cannot be realized.
Confocal microscopes have become a core technology for biology, but have fundamental limitations that hinder their use for neuroscience. The first is slow temporal resolution from raster scanning a laser through the sample to build an image pixel by pixel. Without the ability to parallelize excitation to arbitrary locations within a 3D volume, it is impossible to monitor firing patterns of multiple cells simultaneously. This is critical for mapping connectivity of neural circuits and understanding circuit dynamics.
The second limitation is two-dimensional imaging, which is inappropriate for studies of neural circuits. This restricts studies to a small subset of the neurons and limits the scope of the circuits that neuroscientists are trying to map and understand. The third limitation is confocal microscopy’s coupling of one-photon excitation with a pinhole to block out-of-focus fluorescence emission. This results in low signal from trivially low depths in strongly scattering and absorbing samples, such as neurons within the cortex.
Two-photon microscopy provides submicron lateral and axial excitation confinement without requiring a pinhole, and the longer wavelength simultaneously minimizes scattering. When coupled with spatial light modulators (SLMs), two-photon microscopes are capable of parallelized excitation for photoactivation and volumetric imaging. SLMs can come in a variety of forms, including micromirror arrays and liquid-crystal (LC)-on-silicon modulators.
In a two-photon microscope, the micromirror array is imaged to the sample so that pixels turned on reflect light to neurons for excitation, and pixels turned off reflect light to a block. This allows a simple method to illuminate cell bodies. Micromirror arrays also offer response times on the order of 20 kHz, far surpassing the current response time requirements of neuroscience. However, because the micromirror array is an amplitude modulator as opposed to a phase modulator, it is not possible to generate lens functions for probing activity in a 3D volume or to actively redirect light from pixels that are turned off to desired focal point locations in the sample.
These limitations are overcome through use of LC-SLMs in microscopes. The SLM acts as a programmable lens manipulating the wavefront of the excitation source. In its simplest form, the SLM can be used as a programmable prism, redirecting light to a single focal point with a lateral shift. By adding prism functions together, the SLM can be used to create multiple focal points within a 2D plane. Furthermore, by adding weighting functions and lens functions, the SLM can redirect light to hundreds of focal points with a programmable intensity in a 3D volume (Figure 1).
In two-photon microscopes, LC-SLMs enable multisite 3D scanless excitation for photoactivation2,3,4,5,6, as well as high-speed volumetric imaging to record a volume of circuit activity7. This combination provides neuroscientists with a toolbox for in vivo studies deep within the cortex to better understand the physical structure of neural circuits, the relationship of firing patterns, external stimuli and the resulting behavior, and how these processes are altered in the presence of neurological disease.
3D photoactivation
Traditional two-photon microscopes contain galvanometer-scanning mirrors used to raster scan the laser focus through the sample. The mirrors are conjugate to the back focal plane of the objective. The SLM is added to the system through an additional relay prior to the galvanometer scanning mirrors (Figure 2). The addition of the SLM and two lenses transforms the function of the microscope so that it can deliver light to any location in the field of view and simultaneously excite multiple 3D sites and use a fast camera to capture their responses8.
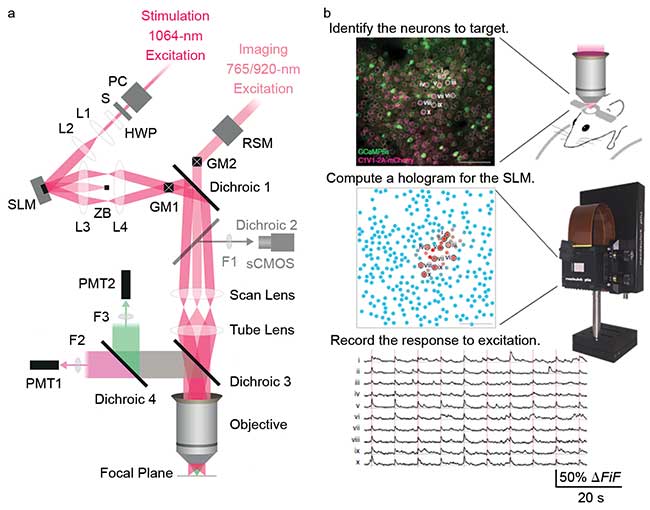
Figure 2. Optical layout of a two-photon microscope with an SLM to enable 3D photoactivation (a). Traditional scanning is used to map the locations of neurons in the sample (b, top). After the cell bodies have been found, specific cells in the field of view can be targeted using the SLM (b, middle). As the cells are excited, the response of the cell bodies can be recorded to map connectivity and record circuit dynamics (b, bottom). Courtesy of A. Packer, L. Russell, H. Dalgleish and M. Hausser, University College London.
In a typical experiment, the galvanometer mirrors raster scan the sample to find the location of cell bodies in the field of view. Holograms then are generated to modulate the wavefront of the source to illuminate individual neurons. This can be used to photoactivate specific cells to replicate firing patterns that have been identified or to manipulate firing patterns that have been observed. Following photoactivation, the response of the surrounding cells can be monitored to understand the impact of photoactivation on the response of the circuit.
When designing the microscope, there are several key criteria that should be considered. The resolution of the SLM determines the number of locations where light can be directed in the sample. The resolution and pixel pitch together determine the dimensions of the volume within the sample that the SLM can excite9. Ideally, the SLM will have a small pixel pitch with high resolution so it can steer to wide angles without under-filling the objective and sacrificing the lateral and axial excitation confinement.
The temporal phase stability of the SLM also is important to ensure reliable excitation. This is particularly important when dividing the light among many neurons and operating near the minimum threshold for excitation. Finally, the response time of the SLM will have significant impact on replicating the spatiotemporal dynamics, which can occur at rates up to 1 kHz110,11.
Volumetric imaging
The ability to manipulate firing patterns is critical to understanding circuit activity, but equally important is the ability to record the response of surrounding neurons at the highest possible frame rate. Traditional two-photon imaging systems build an image volume by mechanically scanning the objective and collecting 2D images (Figure 3). The time required to image the volume can be on the scale of minutes, which is sufficient for static samples. In neuroscience, the dwell time requirement coupled with indicators with limited brightness results in the inability of traditional two-photon imaging to monitor action potentials in complete neural circuits. This opens up the possibility of misinterpretation of action potentials because of the interaction of localized excitation with animal movement.
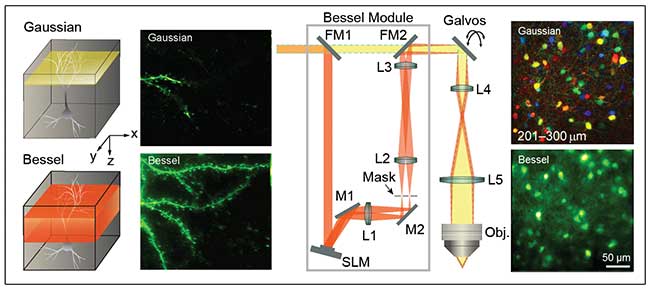
Figure 3. Comparison of Gaussian and Bessel imaging of a mouse dendritic spine (left). Scanning of a Gaussian focus coupled with dwell time requirements for fluorescence excitation leaves a small portion of the sample illuminated and an increased likelihood of activity occurring without fluorescence excitation. Bessel imaging monitors a volume at the same rate of 2D imaging with Gaussian illumination. The “Bessel module” easily integrates with existing 2P microscopes without software changes, enabling easy adaptation of existing microscopes and significantly enhanced capability (middle). A demonstration of the Bessel module used for imaging inhibitory neurons in a mouse. With Gaussian imaging, a series of 2D scans are required to build the 3D projection, but the Bessel module enables imaging the entire volume without axial scanning (right). Courtesy of Na Ji, Janelia Research Campus.
One solution for high-speed volumetric imaging, presented by Na Ji, group leader at the Janelia Research Campus of the Howard Hughes Medical Institute, uses a Bessel focus-scanning technology (BEST) that samples activity in a volume with hundreds of microns in each dimension in the equivalent time that a Gaussian two-photon microscope images a 2D plane6.
The module for 3D imaging is simple and widely compatible with existing microscopes, consisting of an SLM, a static amplitude mask and three lenses (Figure 3). The lenses relay the image of the SLM to the sample. The amplitude mask is a static patterned mirror that selectively transmits the first diffracted order. The optional flip mirrors at the entrance and exit of the module allow optical addition or removal of volumetric imaging so that structures can be imaged with traditional Gaussian illumination if desired. The use of SLMs here allows flexible generation of Bessel foci of varying lateral sizes, axial lengths and axial intensity distribution, permitting users to optimize BEST for specific samples.
Ji has demonstrated the approach for enabling discoveries for neurobiology by imaging the calcium dynamics of volumes of neurons and synapses in fruit flies, zebrafish larvae, mice and ferrets in vivo. Calcium signals in objects as small as dendritic spines could be resolved at video rates. High-speed volumetric imaging is a critical advancement for microscopes adapted specifically to the needs of the neuroscience community.
The combination of SLMs, two-photon microscopy, calcium imaging and photoactivation is leading to advanced tools for neuroscientists to monitor and manipulate the activity of neural circuits in the brain. The methods require minor modifications to existing microscopes, allowing researchers to inexpensively and readily adapt existing tools to support 3D photoactivation with high-speed volumetric imaging. This significantly enhances capabilities of microscopes, providing a complete tool enabling studies of neural circuits, expanding the field of view, the depth and the temporal limits at which neuroscientists can monitor and manipulate circuit activity.
Meet the author
Anna Linnenberger is a senior scientist at Meadowlark Optics. Her education includes a B.S. in computer engineering and an M.S. and Ph.D. in electrical engineering. She has spent the last 16 years supporting and advancing the Meadowlark Optics reflective SLM product line. Her research interests include optical tweezing, optogenetics, tissue engineering and biological imaging; email: [email protected].
References
1. R. Yuste (2011). Imaging. Cold Spring Harbor, N.Y.: Cold Spring Harbor Press.
2. B.O. Watson et al. (2009). Two-photon imaging with diffractive optical elements. Front Neural Circuits, 3:6. Epub 2009/07/29.doi: 10.3389/neuro.04.006.2009. PubMed PMID: 19636390.
3. V. Nikolenko et al. (2008). SLM microscopy: Scanless two-photon imaging and photostimulation with spatial light modulators. Front Neural Circuits, Vol. 2, Issue 5. Epub 2009/01/09. doi: 10.3389/neuro.04.005.2008. PubMed PMID: 19129923; PubMed Central PMCID: PMC2614319.
4. C. Lutz et al. (2008). Holographic photolysis of caged neurotransmitters. Nat Methods, Vol. 5, pp. 821-827.
5. A.M. Packer et al. (2015). Simultaneous all-optical manipulation and recording of neural circuit activity with cellular resolution in vivo. Nat Methods, Vol. 12, pp. 140-146.
6. E. Valentina et al. (2015). All-optical interrogation of neural circuits. J Neurosci, Vol. 35, Issue 41, pp. 13917-13926.
7. R. Lu et al. (2016). Video-rate volumetric functional imaging of the brain at synaptic resolution. bioRxiv: 058495.
8. B.O. Watson et al. (2010). Two-photon microscopy with diffractive optical elements and spatial light modulators. Front Neurosci, 4:29. Epub 2010/09/23. doi: 10.3389/fnins.2010.00029. PubMed PMID: 20859526; PubMed Central PMCID: PMC2940544.
9. W. Yang et al. (2016). Simultaneous multi-plane imaging of neural circuits. Neuron, Vol. 89, Issue 2, pp. 269-284.
10. E. Ronzitti et al. (2017). Sub-millisecond optogenetic control of neuronal firing with two-photon holographic photoactivation of Chronos. Nat Neurosci, Vol. 20, pp. 620-628.
11. R.H.R. Hahnloser et al. (2002). An ultra-sparse code underlies the generation of neural sequences in a songbird. Nature, Vol. 419, Issue 6902, pp. 65-70.
/Buyers_Guide/Meadowlark_Optics_Inc/c9201
Published: September 2017
Glossary
- volumetric imaging
- Volumetric imaging refers to the capture, visualization, and analysis of three-dimensional (3D) information from a volume of space. Unlike traditional two-dimensional (2D) imaging, which provides information along a single plane, volumetric imaging captures data throughout a volume, enabling the reconstruction of a 3D representation.
Key features and applications of volumetric imaging include:
3D data acquisition: Volumetric imaging techniques acquire data from multiple perspectives or...
FeaturesMicroscopyBiophotonicsAnna LinnenbergerMeadowlark Opticsvolumetric imaging