With its swift data acquisition, computational imaging is expanding diagnostics and bringing scientists into a whole new realm of biomedical research.
By Hank Hogan
A picture may be worth more than a thousand words after passing through mathematical algorithms and modern computer hardware. Such a process, known as computational imaging, can speed up data acquisition significantly, thereby allowing the use of diagnostics for a wider population of patients. It enables researchers to study new biomedical processes, using less expensive equipment. And clinicians can diagnose disease using deep learning, a form of artificial intelligence (AI).
Biomedical computational imaging systems form, reconstruct, and/or enhance images created from photon-based sources, x-rays, or magnetic resonance (MR), for instance. Improving imaging can be a formidable task, however.
“It is an emerging interdisciplinary field and represents the integration between science, technology, engineering, mathematics, and computer science,” said Behrouz Shabestari, program director for optical imaging and spectroscopy at the National Institute of Biomedical Imaging and Bioengineering (NBIB) in Bethesda, Md. “Everything is involved in there. That is the major advantage, or the challenge.”
‘[Biomedical computational imaging] represents the integration between science, technology, engineering, mathematics, and computer science.’ — Behrouz Shabestari, NBIB
Compressed sensing is a computational imaging method that offers the significant advantage of reducing patient exposure to ionizing radiation, because fewer x-ray photons are used than with conventional sensing methods. With this approach, only important data is captured. Think of a butterfly against a featureless white background. Most of the scene can be eliminated, since it contains no butterfly-related information. This is how compressed sensing confers benefits.
“Because it [generates] less data, it’s good for storage, archiving, and transfer,” Shabestari said. “But high-quality images can be reconstructed, and at a significantly increased speed.”
Siemens Healthineers, which has its North American headquarters in Malvern, Pa., uses compressed sensing in some of its MR products. The company was the first to be cleared to do so by the FDA in 2017. According to Wesley Gilson, 3 Tesla product manager for the MR business at Siemens Healthineers, the amount of
image speedup depends upon the nature of the data. Compressed sensing has
less of an impact on a 2D static study, for instance, than on capturing the 2D dynamics of a beating heart in a movie-like fashion. This process uses 2D plus time, which results in a 10- to 12-fold imaging acceleration over noncompressed sensing methods. Even more speed is possible, according to Gilson, when imaging the liver. Here, the process is more complex and involves imaging in 3D plus a time component.
“We’re able to realize up to 40-fold acceleration,” Gilson said of using 3D plus time.
Imaging procedures of the heart and liver can require patients to repeatedly hold their breath to minimize the blur caused by chest movement. The process must be repeated as many as 20 times in the case of the heart — a challenge for those with cardiac disease. Faster data acquisition improves patient compliance and expands the size of the population that can be imaged.
Because these diagnostic applications are being used clinically, computational imaging needs to be fast. Siemens Healthineers has found ways to determine the correct sampling rate for the application by using optimized algorithms and taking advantage of computer hardware advancements, according to Gilson. Powerful, specialized GPUs — such as those found in high-end gaming and cryptocurrency systems, for example — are designed and engineered to handle highly parallel operations, which are also common in computational imaging.
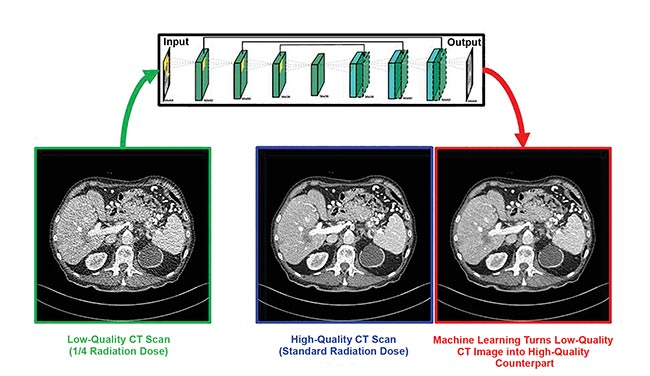
Biomedical computational imaging uses lower doses of ionizing radiation while still producing high-quality images. Deep learning makes quarter-dose CT images up to image quality at normal CT dose. CT denoising neural network by Rensselaer Polytechnic Institute, Sichuan University, and Harvard University. Courtesy of Ge Wang/Rensselaer Polytechnic Institute.
Before these optimizations were possible, reconstructing data into an image using 3D plus time took hours, Gilson said. Now it is done so quickly that it no longer requires long waits and thereby does not impact clinical use.
Computational imaging is also benefiting the optical domain. Jinyang Liang, an assistant professor of optics at the Montreal-based Institut National de la Recherche Scientifique, discusses a compressed sensing technique in a March study1 published in Optics Letters.
“We have developed a new photography approach that allows conventional CCD or CMOS cameras to have ultrahigh imaging speeds of more than a million frames per second,” he said.
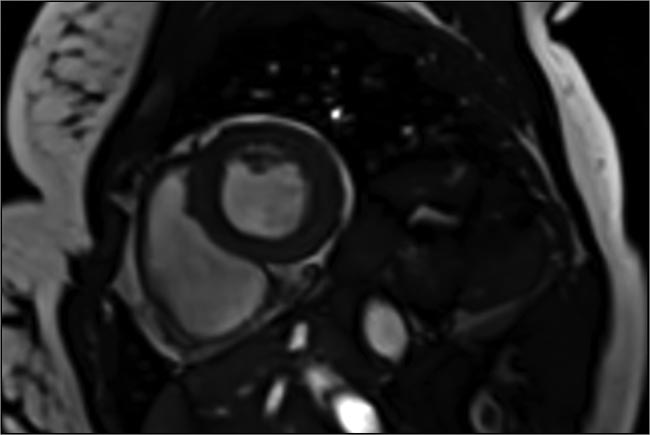
A cardiac image acquired in a single heartbeat, using compressed sensing to speed up image capture 10 to 12×. Courtesy of Siemens Healthineers.
The researchers achieved this speed by combining a digital micromirror device with a synchronized galvanometer scanner. They used the combination to encode a transient image in space and shear it in time. A standard camera captured the resulting shots, with the single images stitched together into a high-speed movie.
Five years ago, it would have taken hours to produce a single image, but with computational imaging this can now be accomplished in real time.
The new imaging technique used in the study has two biomedical applications. The first involves phosphorescent nanoparticles, which have a temperature-dependent emission lifetime. The ability to simultaneously measure this lifetime at many locations in tissue provides researchers with a nanothermometer that could be useful in photodynamic therapy studies.
The second application is neuroimaging. Optically tracing the dynamics of the membrane voltage of neurons requires high sensitivity, along with the ability to record at high speed as an event occurs. According to Liang, their technique offers these capabilities if they use the right camera — such as an electron-multiplying CCD or scientific CMOS camera. His group is now conducting research in this area.
Another optical computational imaging technique involves full-field OCT of the eye, performed without dyes, genetically encoded proteins, or any other biochemical labels. In a collaborative study (published in Biomedical Optics Express)2, lead author Jules Scholler, a graduate student at the Langevin Institute in Paris, noted that their technique involves combining static and dynamic OCT contrast with optional fluorescence contrast for validation, along with computing and applying corrections to the captured signal in real time.
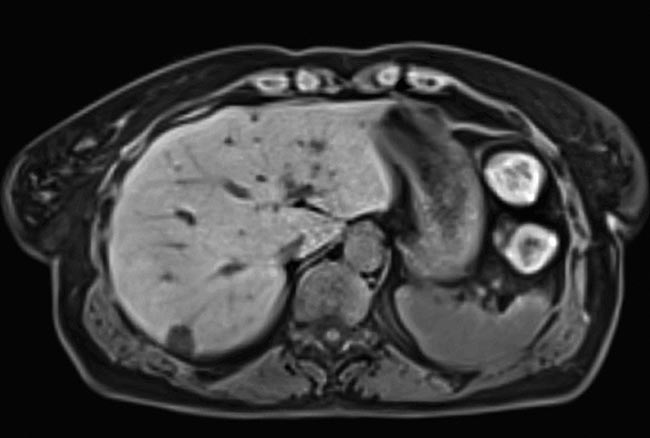
An image of the liver of a free-breathing patient, taken with compressed sensing. Data acquisition accelerated as much as 40-fold. Courtesy of Siemens Healthineers.
The setup allows imaging within the eye on timescales that range from milliseconds to hours. It can probe subcellular biological activity within bulk 3D tissue. To achieve this, the scientists took advantage of advancements in the processor and improvements in bus technology, both of which increased the computational imaging speed substantially.
Five years ago, it would have taken hours to produce a single image, but this can now be accomplished in real time. The group achieved these results in stationary tissue samples, according to Scholler. Imaging at this fine level of spatial resolution in a living eye, however, is more difficult.
“We still need to correct for eye motion, which is a challenge because it requires real-time tracking and compensation with a micrometer accuracy,” Scholler said. “This is higher accuracy than that required by current clinical ophthalmic imaging techniques because our resolution is higher.”
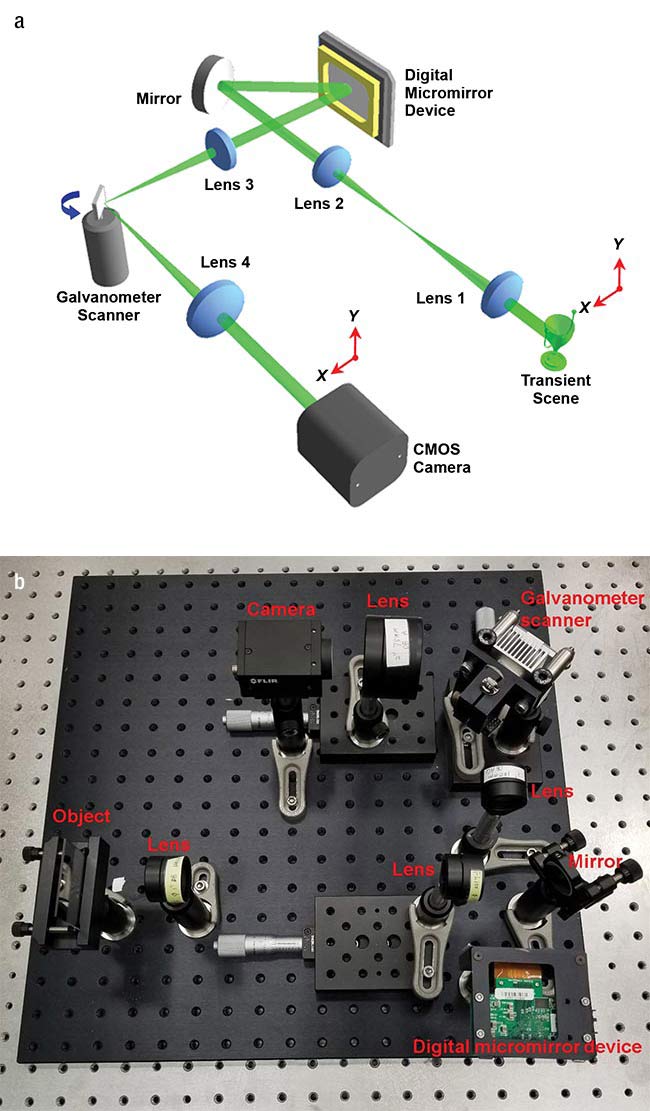
Researchers used a digital micromirror device and a galvanometer scanner — schematic (a) and actual layout (b) — along with computational imaging to transform a standard camera into one capable of a speed of a million frames per second. Courtesy of Jinyang Liang/Institut National de la Recherche
Scientifique.
Using modern, advanced GPUs should make this possible and would permit live-eye imaging microscopy in 3D samples, said Kate Grieve, senior author of the study and a researcher at the Quinze-Vingts National Ophthalmology Hospital in Paris.
“Detecting cell dynamics allows us to access functional measurement on the cellular scale and detect malfunctioning cells,” she said, “which has potential to improve diagnostic precision and disease understanding, and suggest avenues for therapy.”
Implementing this capability requires good hardware, Grieve added. The current setup uses a sensor that was custom developed for this low signal-to-noise application. The optical design must also be of a high enough quality to enable optimization of the image and efficient post-processing computation.
One possible use for this system is in early disease detection, which may be accomplished using AI diagnostics. According to Aydogan Ozcan — a professor in electrical and computer engineering at the University of California, Los Angeles — deep learning is attracting interest for image reconstruction and enhancement.
In deep learning, a neural net learns to classify items by first working through a training set. As it does so, the system adjusts the relative weight of its artificial neurons until it successfully categorizes items in the sample set.
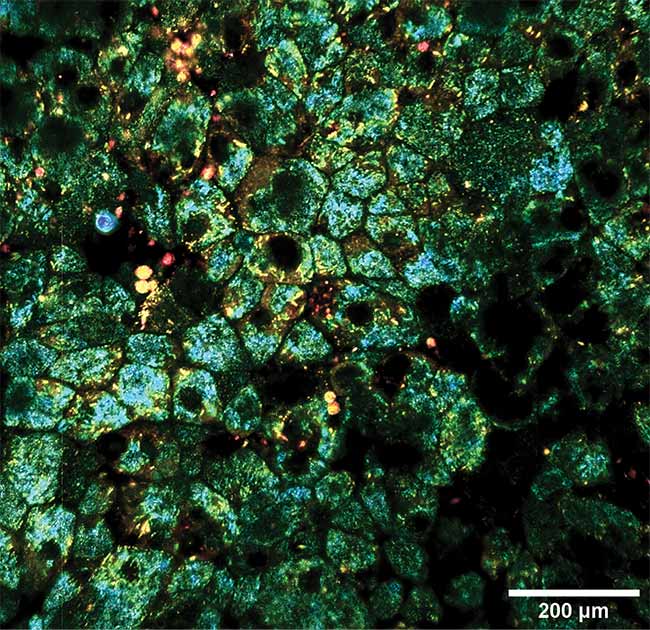
Dynamic and static contrast OCT of the eye, taken with a computational imaging technique that can image whole tissue at a subcellular level for hours at a time. Courtesy of Kate Grieve/Quinze-Vingts National Ophthalmology Hospital.
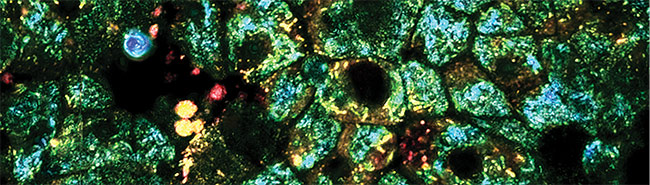
Ozcan’s team used deep learning techniques to create virtually stained microscopic images. They took unlabeled tissue and captured its autofluorescence image — the one arising from normally occurring fluorescence in the tissue. The researchers then trained the neural network by giving it thousands of autofluorescence images, as well as the corresponding ones of the same tissue after staining.
A study published in the March issue of Nature Biomedical Engineering3 describes how, when using this approach, the system learns to transform an autofluorescence image into one that shows what the tissue would look like if it were chemically stained. Bypassing the traditional staining process saves time, money, and lab space, while preserving tissue for later analysis by other means, Ozcan said. This type of computational imaging could be applied to other forms of microscopic examination that today depend upon the use of contrast agents. Thus, it could be used in surgery rooms or in telepathology for rapid diagnosis.
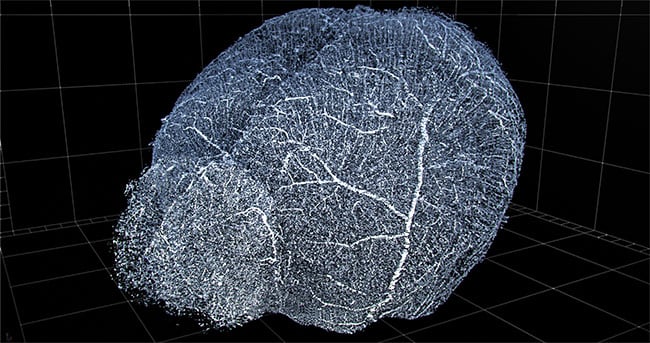
Computational imaging and automation enable 3D reconstruction of vasculature for viewing and extraction of statistics. Shown here is a 3D reconstruction of mouse brain vasculature. Courtesy of 3Scan.
“Beyond its clinical applications, this method could broadly benefit the histology field and its application in life science research and education,” Ozcan said.
AI and deep learning could become increasingly important to computational imaging in the future. For instance, San Francisco-based 3Scan currently makes an automated 3D histopathology system. The robot sections tissue into thin slices, images them with a line-scan camera, and then stacks and stitches the 2D slice data into a 3D reconstruction of the original tissue. Users can then seamlessly move through the 3D representation, zooming in as desired.
“You can get down to a cellular-scale resolution, but [capture] whole-organ volumes for research animals. It’s kind of a cross between CT and slides,” said Megan Klimen, chief operating officer and co-founder of 3Scan. “The large value proposition in the future is to be able to do more deep learning for different types of data sets.”
The company plans to develop new products that will provide statistical analysis of important biological parameters, such as density and other data about the vasculature system. Such analysis should eventually significantly benefit from deep learning, an indication of the direction in which the entire field of computational biomedical imaging is heading.
References
1. Xianglei Liu et al. (2019). Single-shot compressed optical-streaking ultra-high-speed photography. Opt Lett, Vol. 44, Issue 6, pp. 1387-1390.
2. J. Scholler et al. (2019). Probing dynamic processes in the eye at multiple spatial and temporal scales with multimodal full field OCT. Biomed Opt Express, Vol. 10, Issue 2, pp. 731-746.
3. Y. Rivenson et al. (2019). Virtual histological staining of unlabelled tissue-autofluorescence images via deep learning. Nat Biomed Eng, https://doi.org/10.1038/s41551-019-0362-y.