Collaborators from Carnegie Mellon University, the University of Pittsburgh, and Brown University have described a microscopy technique and set of protocols that overcome a bottleneck to the expansion microscopy method. The collaborators developed “Magnify” as a variant of expansion microscopy that uses a hydrogel that retains a spectrum of biomolecules, offers a broader application to a variety of tissues, and increases the expansion up to 11× times linearly or approximately 1300 folds of the original volume.
Through the expansion microscopy process, samples are embedded in a swellable hydrogel that homogenously expands to increase the distance between molecules, which allows them to be observed in greater resolution. This allows nanoscale biological structures that previously could be viewed only via expensive high-resolution imaging techniques to be seen with standard microscopy tools.
In addition, the researchers said in their paper, “Current expansion microscopy protocols require prior treatment with reactive anchoring chemicals to link specific labels and biomolecule classes to the gel.”
In developing the Magnify method, according to the researchers, the team developed a gel that was mechanically sturdy to retain nucleic acids, proteins, and lipids without the need for a separate anchoring step.
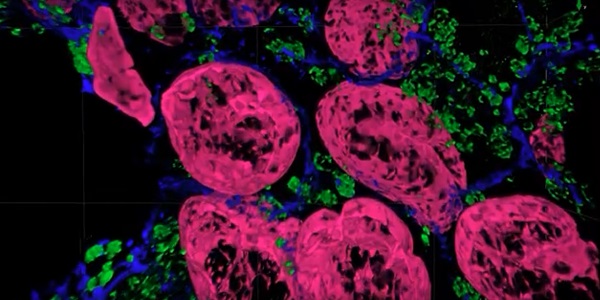
A tissue sample imaged with the Magnify expansion microscopy method. Courtesy of Carnegie Mellon University.
Yongxin (Leon) Zhao, the Eberly Family Career Development Associate Professor of Biological Sciences at Carnegie Mellon, said, “We overcame some of the long-standing challenges of expansion microscopy. One of the main selling points for Magnify is the universal strategy to keep the tissue’s biomolecules, including proteins, nucleic acids, and carbohydrates, within the expanded sample.”
Keeping different biological components intact matters, since previous protocols required eliminating many various biomolecules that held tissues together, Zhao said. However, these molecules could contain valuable information for researchers.
“In the past, to make cells really expandable, you need to use enzymes to digest proteins, so in the end, you had an empty gel with labels that indicate the location of the protein of interest,” he said.

Using the Magnify method, the molecules are kept intact, and multiple types of biomolecules can be labeled in a single sample.
“Before it was like having single-choice questions. If you want to label proteins, that would be the version one protocol. If you want to label nuclei, then that would be a different version,” Zhao said. “If you wanted to do simultaneous imaging, it was difficult. Now with Magnify, you can pick multiple items to label, such as proteins, lipids, and carbohydrates, and image them together.”
Co-first author and postdoctoral researcher Aleksandra Klimas said that in addition to its high-resolution imaging properties, the newly described approach is advantageous because of its broad applicability. “Traditionally, you need expensive equipment and specific reagents and training. However, this method is broadly applicable to many types of sample preparations and can be viewed with standard microscopes that you would have in a biology laboratory,” she said.
Doctoral student Brendan Gallagher, an additional co-first author of the work, said that the team tried to make the protocols involved in its method as compatible as possible for researchers who could benefit from adopting Magnify. As a result, Gallagher said, Magnify works with different tissue types, fixation methods, and tissue that has been preserved and stored.
The developed protocols aim to provide a framework for those in neuroscience, pathology, and other biological and medical fields. According to the researchers, the small sizes of monomers, as well as the fast rate of diffusion, mean that Magnify may have applicability to thick tissues and whole organisms. “Magnify would be readily adaptable to generating nanoscale whole organ data sets, which currently rely on either lower-resolution tissue-clearing methods,” they said in their paper.
In addition, they said, because Magnify is a chemical strategy that does not rely on complex optics, its framework can be adapted to a range of imaging modalities and gel chemistries, as well as with other expansion microscopy strategies — which have previously demonstrated compatibility with existing superresolution techniques.
The work was published in Nature Biotechnology (www.doi.org/10.1038/s41587-022-01546-1).