The technique holds the promise of greater accuracy and efficiency for neuroscience, vascular research, and cytopathology.
GARY GREENBERG, HNU-EDGE LLC
Stereo 3D microscopes produce real-time 3D images, but they are usually limited to low-magnification applications, such as dissection. Most compound light microscopes produce flat, 2D images because high-magnification microscope lenses have inherently shallow depth of field, rendering most of the image out of focus.
Because of the shallow depth of focus of microscopes, microscopists usually prepare 5- to 10-µm sections of their samples to eliminate the problem of out-of-focus blur. This is problematic because the average cell is between 10 and 30 µm. Examining 5- to 10-µm sections leads to undersampling errors, resulting in misinterpretation of the data.
Most people’s time on microscopes is spent searching for appropriate fields of view from which to acquire images. More information is gleaned in a shorter period of time when viewing in stereo because 3D provides a clearer understanding of the structures within a specimen compared to 2D viewing.
High-power, real-time 3D viewing could provide a significant benefit for investigations that employ the micro-manipulation of probes to stimulate, record, or move cells from one place to another. Real-time 3D viewing also would be highly advantageous for in situ investigations of hard tissues, such as bones, teeth, and shells.
HNu-Edge LLC has introduced a patented, high-power 3D microscope with extended depth of focus that functions in real time. Proprietary Panfocal software provides 3D reconstruction, 3D display, and 3D measurement in samples from 20 to >200 µm thick. This 3D Panfocal approach provides a powerful alternative to conventional 2D microscopes and offers a cost-effective substitute to expensive confocal microscopes.
The first high-power 3D microscope was invented by John Riddell more than 160 years ago1. Riddell placed a prism behind the objective lens, directing the image from the left half of the objective to the left eye and the right half of the objective to the right eye. The result was pseudostereo — in other words, the depth information was reversed, making interpretation of the image difficult.
Since the turn of the 21st century, confocal microscopes have popularized high-magnification reconstruction 3D microscopy. It was first patented by Marvin Minsky in 19572. Minsky did not develop the idea into a product and subsequently co-founded the Massachusetts Institute of Technology’s Artificial Intelligence Lab.
In the 1960s, Czechoslovak scientist Mojmír Petrán developed the tandem scanning confocal microscope, which uses a spinning disk with multiple pinholes to simultaneously illuminate and examine the entire selected area of the specimen3. Petrán’s apparatus allowed for multiple focal levels of the specimen to be collected quickly. The resulting stack of images could then be viewed as a reconstructed image, dramatically increasing depth of focus and producing a fully focused image of a thick specimen.
The laser scanning confocal microscope was developed by David Egger and Paul Davidovits, and they published a paper on their findings in 19694. It improved signal strength and specificity, making it particularly useful for fluorescence imaging. Newer methods, such as multiphoton and light-sheet microscopy, have extended the fluorescence capabilities of these sophisticated 3D microscopes.
The Edge 3D produces full-focus 3D images by using automated z-stacking that acquires a series of images from successive focus levels through thick specimens. Panfocal software removes the out-of-focus portions of each image in the stack and reconstructs a full-focus 2D image that represents the 3D volume of the data.
The system is fully integrated and automated. It captures the images and then processes, analyzes, and measures them in 3D. The depth information is displayed in several ways (Figure 1), including compressed 2D images of the 3D volume, stereo 3D images, calibrated color depth images, and 3D motion parallax video loops that reveal the depth information without 3D glasses.
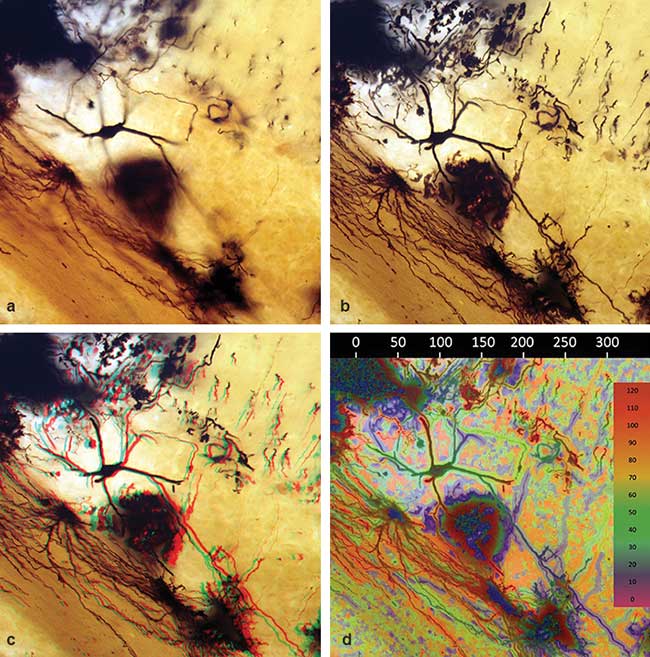
Figure 1. Golgi-stained, 120-µm section of neurons from rat cerebellum imaged with a 40× objective lens: a conventional 2D optical section (a), a compressed 3D image reconstructed from multiple optical sections (b), a red/cyan stereo 3D image of the data set (c), and an X, Y, and Z color depth map calibrated in micrometers (d).
High-magnification Panfocal 3D microscopes offer several advantages. Increased depth of focus provides more accurate information by displaying hidden depth information. The ability to examine fully focused thick samples can reveal interrelationships between structures within the specimen that might otherwise go unnoticed. The microscopes increase productivity, reduce misdiagnoses, and improve comprehension of the data.
The technology provides 3D imaging of thick samples without the need for expensive confocal equipment. The use of computer algorithms to remove the out-of-focus portions of a conventional wide-field image allows for a wider range of optical imaging modes, such as 3D phase contrast, 3D bright-field and dark-field, both reflected and transmitted illumination, and epifluorescence.
This means that 3D imaging methods can be applied to tissues using nonfluorescence staining techniques, such as Golgi stains and immune histochemical methods. The advantage of these nonfluorescence methods is the achievability of the specimens. This permits detailed image capture and analysis over long periods, whereas fluorescence methods require fast and immediate capture of images before the signal quenches and disappears.
The best application of 3D imaging is in samples that are thicker than the standard 5-µm section. In fact, many samples do not need to be sectioned and can be investigated as in situ preparations, including dissected whole-mounts such as mesentery or retina. Stereo viewing is a powerful tool for understanding the interrelationships between structures within a thick specimen.
The additional depth information can reduce misdiagnoses and increase productivity in a wide range of applications, including neuroscience, vascular research, cytopathology, embryology, 3D tissue culture, entomology, and plant biology.
Neuroscience. Understanding the nervous system is a fundamentally 3D problem. Nerve cells span relatively large distances and connect with one another in 3D patterns. Because multifunctional, nonconfocal 3D microscopes have the capacity to investigate nonfluorescence specimens, these preparations are archival compared to fluorescent specimens, which fade and quench very rapidly.
In fact, a selected field of view usually fades within minutes of viewing, whereas nonfluorescence specimens can be investigated for long periods, allowing more complete information to be gleaned from each specimen. Significantly more information is revealed with 3D images compared to conventional 2D images.
Vascular Research. The interconnectedness of blood vessels makes vascular research an important application for 3D microscope analysis. Three-dimensional presentation of the depth information can improve the science by revealing interrelationships among structures.
Cytopathology. This application represents a perfect example of how 3D microscopes can increase productivity. The conventional standard is to cut a biopsy into multiple serial sections, each 5 µm thick. The cytopathologist examines these sections to attain an understanding of the cells and tissues within the 3D biopsy. Significant time and energy can be saved by cutting 50-µm sections for use with a 3D microscope. The cytopathologist need examine only one-tenth the number of sections, with the added benefit of seeing the cells and tissues in relationship to one another in 3D. It also has been demonstrated that the use of 3D microscopes can significantly reduce false diagnoses of atypical squamous cells in Pap smear tests5.
Embryology. The development of cells and tissues depends upon the local environment in which the cells reside. Cell-cell and cell-extracellular matrix interactions guide development of the organism. Embryological research is often performed using thick tissue preparations, making it an excellent application for nonconfocal 3D microscopes.

Figure 2. The claw of the fungus gnat Bradysia difformis (Diptera: Sciaridae) imaged with a 20× objective lens: a conventional 2D optical section (a), a compressed 3D image reconstructed from multiple optical sections (b), and a red/cyan stereo 3D image of the data set (c).
3D Tissue Culture. Recently, 3D cell culture systems have become more relevant. Compared to 2D culture systems, 3D cell cultures more closely represent the microenvironments that occur in vivo. The multifunctional nature of Panfocal 3D microscopes provides a valuable tool for visualizing 3D cell cultures. The culture can be seen using fluorescent markers or without markers via phase contrast or oblique illumination.
Entomology. 3D microscopes are an excellent tool for visualizing insects in high resolution with full depth perception (Figure 2).

Figure 3. Stamen with pollen from a water hyacinth flower in the foreground and trichomes in the back-ground, imaged with a 10× objective lens: a conventional 2D optical section (a), a compressed 3D image reconstructed from multiple optical sections (b), and a red/cyan stereo 3D image of the data set (c).
Plant Biology. Plants form intricate 3D structures that can be appreciated only when seen as an unsectioned whole specimen (Figure 3).
Meet the author
Gary Greenberg is COO of HNu-Edge LLC, a company that innovates 3D imaging systems. He received his Ph.D. in developmental biology from University College London, after which he became an assistant professor at the University of Southern California. He is working with HNU Photonics on a multifunctional microscope for the International Space Station; email: [email protected].
References
1. J.L. Riddell (1854). On the binocular microscope. J Cell Sci, s1-2, pp.18-24.
2. M. Minsky (1957). Patent US3013467A, Microscopy Apparatus.
3. M. Egger and M. Petrán (1967). New reflected-light microscope for viewing unstained brain and ganglion cells. Science, Vol. 157, pp. 305-307.
4. P. Davidovits and M.D. Egger (1969). Scanning laser microscope. Nature, Vol. 223, p. 831.
5. R. Ramsamooj et al. (2002). Real-time, high-definition, three-dimensional microscopy for evaluating problematic cervical Pap smears classified as atypical squamous cells of undetermined significance. Cancer Cytopathol, Vol. 96, Issue 3, pp. 181-186.
See www.hnu-edge.com/biophotonics.html for a motion 3D video loop of the images in this article that reveals the depth information without 3D glasses.