A fluorescently labeled nucleic acid instrument with a stem-loop structure helps to identify a specific cancer microRNA sequence.
SULAYMAN A. OLADEPO, KING FAHD UNIVERSITY OF PETROLEUM AND MINERALS
A smart probe is a hairpin oligonucleotide, essentially a string of nucleic acids, that is labeled with a fluorophore on one end and tethered with successive guanosine residues on the other end1. Extrinsic dye labels with suitable absorption are typically used for fluorescence quenching, which signals a reaction. But recent research has shown that guanosine moieties can provide effective fluorescence quenching for common fluorescent dyes due to guanosine’s low oxidation potential2. Thus, guanosine is used as a quencher in smart probes, which can detect biomarkers of cancer and other conditions.

Courtesy of iStock.com/mkarco.
Several methods of detection have been used for microRNAs in past research. These include polymerase chain reaction, northern blotting, in situ hybridization, and microarray methods. While these traditional methods are still in use, they present certain challenges that make their continued adoption less attractive.
For instance, the selectivity and specificity of polymerase chain reaction methods are not in question; however, the detection process takes a long time because it requires purified isolated RNA, and the process is highly involved. Northern blotting is semi-quantitative and cumbersome, and it has low sensitivity and requires a large amount of sample. The high throughput capability of microarray methods notwithstanding, they present limited sensitivity, selectivity, and specificity.
These limitations have led to the development of new and emerging methods for microRNA detection. Such methods include electrochemical and colorimetric approaches and nucleic acid amplification, as well as the smart probe-based, mix-and-read homogeneous method presented here.
Smart probe makeup
A smart probe is primarily designed to have a stem-loop structure (Figure 1) that is maintained at room temperature. The loop sequence of the probe is perfectly complementary to the target sequence of interest — the subject of research or biomedical examination — while the stem strands are self-complementary and are strongly held together via hydrogen bonding. Thus, at room temperature and in the absence of the target sequence, the probe maintains its hairpin, stem-loop conformation. This conformation places the fluorophore and quencher in close proximity, so the fluorescence signal is effectively reduced.
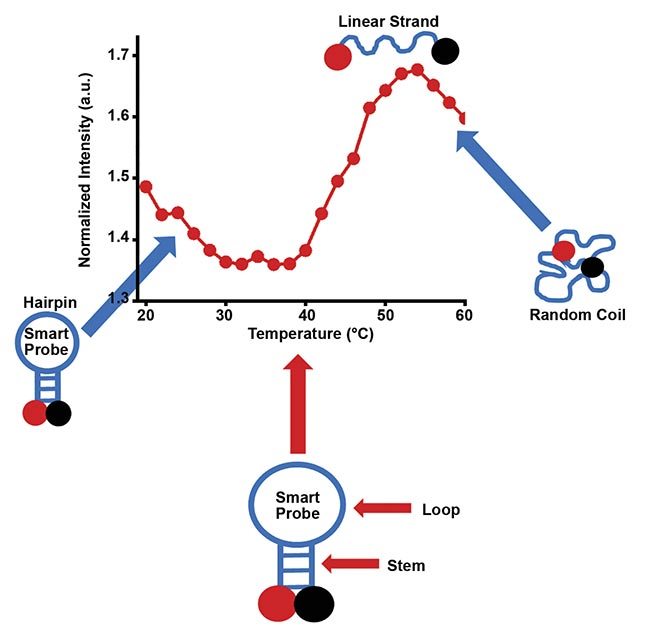
Figure 1. A schematic showing the smart probe in its stem-loop conformation, as well as its various conformations at different temperatures. The temperature profile of its fluorescence signal is also shown for various conformations. Courtesy of King Fahd University of Petroleum and Minerals.
The stem of the smart probe is closed, and the fluorophore and quencher somewhat overlap each other. This conformation is referred to as the off state of the probe. When the target sequence is present, however, the probe spontaneously hybridizes with the target while undergoing a conformational change that forces apart the fluorophore and the guanosine quencher. This duplex formation produces an intense fluorescence signal, since the quencher is now far away from the fluorophore and is therefore unable
to quench the fluorescence signal (on state).
If the target has a single-base mismatch (Figure 2), often associated with a mutation, such that the sequence is not perfectly complementary with the loop of the probe, spontaneous hybridization also happens due to nonspecific interaction between the probe and the mismatch sequence. And the duplex is less stable, which leads to a fluorescence signal that is less than that produced with the perfect target from which maximum fluorescence is recorded. This difference in signal levels shows that the smart probe is able to subtly discriminate between a perfect target and a mismatch sequence. The probe is also very sensitive in detecting low levels of the perfect target sequence. In fact, the constrained structure of this hairpin probe is responsible for its inherent sensitivity3. Since the probe is labeled with a fluorescent dye, the resulting fluorescence signal is measured by a fluorescence spectrometer.

Figure 2. A hairpin smart probe, when the stem is closed and the fluorophore and quencher are in close proximity (off state). The smart probe can form a stable duplex with the perfect target sequence, resulting in a high fluorescence signal. It can also form a less stable duplex with a mismatch target containing a single-base mismatch and therefore produce less signal, depending on the desired effect. Courtesy of King Fahd University of Petroleum and Minerals.
If temperature is gradually increased beyond room temperature, the stem of the smart probe begins to melt and the fluorophore-quencher pair begins to separate, and they move away from each other. This gradual conformational change can be tracked by the fluorescence signal that is recorded as the temperature increases. Accordingly, at room temperature, when the fluorophore and quencher overlap each other, the stem is closed and there is only a limited fluorescence signal due to quenching by guanosine residues (Figure 1).
As the temperature gradually increases, the stem melts and the fluorescence signal gradually begins to increase. The increase in signal continues as the temperature increases, until the stem is fully open and the fluorophore-quencher pair is maximally separated. This maximum separation corresponds to maximum fluorescence signal.
The midpoint of the inflection region between low and maximum signals represents the melting temperature of the stem of the smart probe. If the temperature further increases, the probe transitions into a random coil conformation in which the relative locations of the fluorophore and quencher are random. Hence the fluorescence signal begins to decrease because the fluorophore and quencher are mostly closer to each other. The entire profile traversed by the temperature-dependent fluorescence signal is consistent with the sigmoidal profile expected of a hairpin probe4. The profile may be referred to as a thermal transition or a thermal denaturation profile.
Temperature-dependent fluorescence measurements carried out on the smart probe (SP)-target mixture produce a thermal transition profile that is different from that produced by the probe alone. If the perfect target is present, the probe spontaneously hybridizes with the target and the conformation changes such that a duplex is formed and the fluorophore and quencher are therefore forced apart. The fluorescence intensifies this result from the changing temperature (Figure 3).
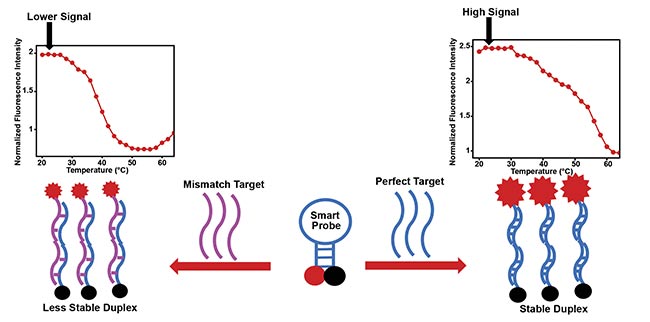
Figure 3. The smart probe and its hybridization with a perfect target form a stable duplex with intense fluorescence signal (right). The smart probe
and its hybridization with a mismatch target form a less stable duplex, resulting in slightly lower fluorescence signal (left). Courtesy of King Fahd
University of Petroleum and Minerals.
Thus, at room temperature a stable duplex is readily formed and produces an intense fluorescence signal. If the temperature is increased, the duplex would gradually melt and the fluorescence signal would gradually decrease. This decrease in signal continues as the temperature increases, until the duplex is completely melted and the probe completely detaches from the target and forms a hairpin. Due to the melting of the stem of the probe, the fluorescence signal may slightly rise again if the temperature is increased beyond this point. If the target contains one base mismatch, the probe also forms a duplex with the mismatch target via nonspecific interactions. This occurs because the entire base sequence is the same as that of the perfect target, except that one single base is different, as shown in the example below:
 However, the duplex would be slightly less stable due to the single-base mismatch, so the signal level produced by this duplex is slightly less than that produced by the duplex of the perfect target. The difference in signal levels between the two duplexes therefore corresponds to whether or not a perfect target is involved in duplex formation. It also serves to show that the smart probe is able to discriminate between the two similar sequences by reporting slightly different fluorescence signal levels.
However, the duplex would be slightly less stable due to the single-base mismatch, so the signal level produced by this duplex is slightly less than that produced by the duplex of the perfect target. The difference in signal levels between the two duplexes therefore corresponds to whether or not a perfect target is involved in duplex formation. It also serves to show that the smart probe is able to discriminate between the two similar sequences by reporting slightly different fluorescence signal levels.
Smart probe target
A smart probe may be used for sequence-specific detection of various targets such as microRNA (miRNA) and other nucleic acid targets. The specific probe described here has been used by researchers at King Fahd University of Petroleum and Minerals for the sequence-specific detection of synthetic miR-21, a cancer biomarker. This synthetic miR-21 sequence was therefore the perfect target sequence used in the work. To prepare for the study, a 100-nanomolar solution of the smart probe consisting of a TE (Tris-EDTA) buffer at pH 7.5 and sodium and magnesium ions was mixed with a 3× excess perfect target and incubated in the dark for ~5 h.
This procedure was followed by temperature-dependent measurements of the fluorescence signal on an FLS920 Fluorescence Spectrometer from Edinburgh Instruments. A thermoelectrically cooled sample holder was used, and real-time temperature control was achieved by means of a TC 125 temperature control unit from Quantum Northwest Inc. To ascertain that the smart probe could sequence-specifically recognize the target and discriminate between it and a similar sequence with a single-base mismatch, a 3× excess mismatch target sequence was also mixed in a 100-nanomolar solution of the smart probe and similarly incubated in the dark for ~5 h before fluorescence was measured. The fluorophore used in the probe was
6-carboxyfluorescein (6-FAM). Excitation and emission wavelengths were respectively 490 and 520 nm, with the emission scan acquired from 500 to
650 nm. An emission maximum of
~520 nm was used for all data analysis.
The temperature-dependent fluorescence signals of the smart probe, SP-target duplex, and SP-mismatch duplex were measured, and the analyzed data was used to generate the thermal transition profiles. The probe performed as expected, generating low signals at low temperatures and high signals at high temperatures when the stem was completely detached.
As for the SP-target duplex, the fluorescence signal was high at room temperature because the probe was able to sequence-specifically recognize this target, hybridize with it, and cause the stem to completely detach and produce the observed high fluorescence signal. The signal decreased as temperature increased because the duplex gradually melted and the fluorophore and quencher gradually approached one another.
A similar profile was recorded for the SP-mismatch duplex, albeit a lower fluorescence signal was generated at room temperature. At room temperature, the fluorescence signal recorded for the mismatch duplex was about 80% of the SP-target duplex. The relatively low fluorescence signal implies that the SP-mismatch duplex is less stable as a result of the single-base mismatch present in the mismatch sequence. This reduced stability is reported by the relative low signal. The signal differential indicates that the smart probe is able to subtly discriminate between the perfect target and the mismatch sequence.
The melting temperature for the smart probe is 63 °C. For the SP-target duplex, it is 47 °C, while for the SP-mismatch duplex it was found to be 37 °C. The wide gaps between the melting temperature values further show that the probe is able to discriminate between the perfect and mismatch target sequences. It also proves that the smart probe meets its design requirements when its melting temperature is at least 5 °C higher than that of the duplex, such that the smart probe must remain a hairpin even after the duplex has completely melted5.
While the subtle discrimination produced by the smart probe at room temperature is acceptable, better discrimination can be obtained at ~52 °C. Although the signal recorded by the SP-target duplex is lower than that recorded at room temperature, both the smart probe and the SP-mismatch duplex have essentially equal signal levels at 52 °C. This shows that the probe can present excellent discrimination at this temperature because the signal generated by the SP-mismatch duplex is the background signal generated by the smart probe alone. Therefore, fluorescence signals were measured at this temperature on separate samples following the usual incubation procedure.
The fluorescence signal generated by the SP-target duplex at 52 °C is far higher than those generated by the smart probe alone and by the SP-mismatch duplex. Thus, the smart probe described here can offer excellent discrimination between the perfect target and mismatch sequences despite the close similarity between the two sequences.
The target oligonucleotides used in this work are synthetic nucleic acids that mimic the miR-21 cancer biomarker. The work laid a strong foundation for adapting the smart probe-based homogeneous miRNA detection method for detecting miR-21 in real samples, such as cancer cell lines. Laboratory experiments involving such in vivo miR-21 detection using this smart probe-based method are currently underway at King Fahd University of Petroleum and Minerals.
Meet the author
Sulayman A. Oladepo is an assistant professor in the Chemistry Department at King Fahd University of Petroleum and Minerals (KFUPM) in Saudi Arabia. His current research interests lie in the development of smart molecular probes for cancer detection, bioanalytical method development, and spectroscopic instrumentation. Prior to joining KFUPM, Oladepo worked on a project involving the development of low-frequency-shift Raman instrumentation for pharmaceutical respirable powders; email: [email protected].
References
1. J.P. Knemeyer et al. (2000). Probes for detection of specific DNA sequences at the single-molecule level. Anal Chem, Vol. 72, pp. 3717-3724.
2. C.A.M. Seidel et al. (1996). Nucleobase-specific quenching of fluorescent dyes. 1. Nucleobase one-electron redox potentials and their correlation with static and dynamic quenching efficiencies. J Phys Chem, Vol. 100, pp. 5541-5553.
3. G. Bonnet et al. (1999). Thermodynamic basis of the enhanced specificity of structured DNA probes. Proc Natl Acad Sci USA,
Vol. 96, pp. 6171-6176.
4. S. Tyagi and F.R. Kramer (1996). Molecular beacons: probes that fluoresce upon hybridization. Nat Biotechnol, Vol. 14,
pp. 303-308.
5. S.A. Oladepo (2018). Design and characterization of a singly labelled fluorescent
smart probe for in vitro detection of miR-21.
Appl Spectrosc, Vol. 72, No. 1, pp. 79-88, www.doi.org/10.1177/0003702817736527.