Already important to the detection and identification of bacteria, surface-enhanced Raman spectroscopy (SERS) is being explored for cancer diagnosis, especially in its early stages.
FRAN ADAR AND MARUDA SHANMUGASUNDARAM, HORIBA INSTRUMENTS INC.
Raman spectroscopy is a spectroscopic technique used to measure vibrational transitions in molecules, yielding detailed information on molecular structures and their environments. Given that it is a light-scattering technique that uses a visible laser for excitation, the spatial resolution (ca. 0.5 λ) is better than 1 µm. While this sensitivity is often not adequate for some applications, the phenomenon called surface-enhanced Raman spectroscopy (SERS) features signal intensities increased by six to 14 orders of magnitude.
While SERS has been known for nearly 40 years, the technique is not universal — it does not work on any arbitrary molecule and is not always reproducible. Given the potential payoff of being able to apply this technology as an analytical tool, researchers have been working to understand the origin of the enhancement. And with this understanding, SERS substrates1 will enable analytical utilization that can be rationally and reproducibly designed.
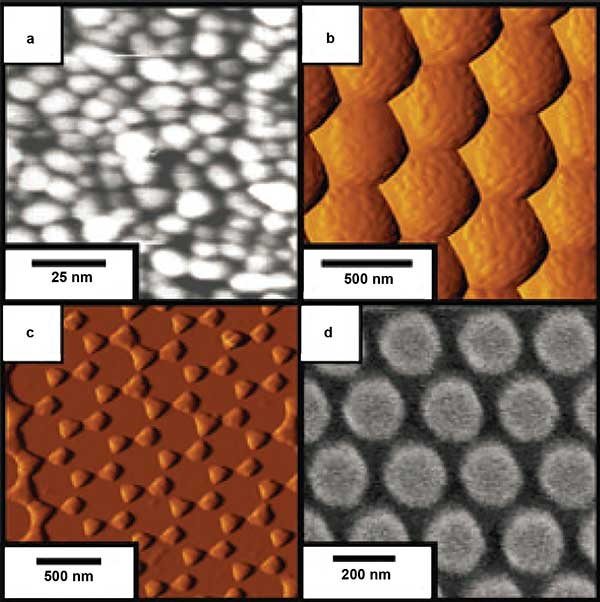

SERS substrates with designs based on plasmonic properties of metals. Metal island film (a), metal film over nanospheres (b), triangular nanoparticle array fabricated with nanosphere lithography (c), and cylindrical nanoparticle array fabricated with electron-beam lithography (d). Reproduced with permission from C.L. Haynes, A.D. McFarland and R.P. Van Duyne (2005). Surface-enhanced Raman spectroscopy. Analytical Chemistry, Vol. 77, pp. 338A-346A.
Chemists have been defining many experimental conditions that are carefully designed to provide analytical information. However, a careful read of their work indicates that they are aware of the limits of reproducibility of the SERS signals from their materials. It is important to note that the original SERS measurements were made on silver electrodes and then with nanoparticles of silver. However, within the past few years it was recognized that the signals were vastly enhanced when an aggregating agent was added2. The role of aggregation was confirmed3 and has led to the evolution of plasmonic theories4. Researchers with a background in physics have produced plasmonic metallic structures optimized for particular laser excitation wavelengths in order to confirm the theoretical underpinning of SERS5.
Types of SERS substrates
SERS-active substrates come in several varieties. They can be “electrodes, island films, well-dispersed nanoparticle sols, nanoparticle films and highly ordered nanostructures,” according to Marek Procházka, a professor at the Charles University in Prague6. The choice of substrate will depend on the purpose of the measurement. For instance, if a researcher wants to measure an analyte in a flowing apparatus, such as a lab-on-a-chip, he or she could use nanoparticles in a solution or deposited on the walls of the glass tubes through which the solution is flowing7. Due to the particles’ semitransparency, it would be possible to observe an analyte adhering to them.
Nanoparticles assembled on a planar support might be of use for detecting a specific analyte by either dipping the support into the relevant solution or depositing a droplet on the support. A planar substrate can also be prepared lithographically, in which case the structures can be accurately engineered to replicate the best enhancing conditions as predicted by the plasmonic theories8. Other approaches include nanosphere lithography or colloidal lithography, and electrochemical deposition of nanowires.
There are also variations on the composition of the nanoparticles. Shell-isolated nanoparticle-enhanced Raman spectroscopy (SHINERS) are spheres of noble metal coated with an ultrathin glass shell. This has the effect of preventing chemical reactions between the metal and the analyte solution, and preventing oxidation of the metal surface, which is of particular concern in the case of silver. In a second scheme, the nanoparticle core can be magnetic and coated with a noble metal film. The advantage of this would be the ability to collect particles magnetically after exposure to the solution of interest.
Characteristics for quantitative SERS substrates include: sensitivity, spot-to-spot uniformity, batch-to-batch reproducibility, stability, the absence of surface contamination so that weakly absorbing analytes can be studied, and the low cost of preparation and ease of use9. For a SERS sensor, sensitivity and selectivity are of ultimate importance. In the words of Procházka10, the process of validation should include “specificity, linearity, accuracy, precision, range limit of detection (LOD) and limit of quantification (LOQ).”
One of the biggest challenges is how to ensure specificity or selectivity. Again, not all molecules are SERS-active, and there may be several SERS-active molecules in a solution competing for the same SERS sites. Given that SERS activity requires the attachment of the analyte molecule to the substrate, one selectivity method is to modify the metal or glass surface with a bifunctional chemical anchor that will capture the analyte of interest. In another scheme, named extrinsic SERS, the analyte is associated with a “Raman reporter molecule” that has an intense and distinguishable Raman signature. This scheme is often used on large biomolecules, such as antibodies11.
Analytical applications of SERS
For analytical purposes, the integration of SERS detection with lab-on-a-chip technology12,13 can provide a paradigm shift in the analytical use of Raman spectroscopy. SERS has been used to detect various analytes in body fluids, such as saliva, urine and blood or serum. Examples include pharmaceuticals such as anti-cancer drugs, narcotics, controlled substances and their metabolites, food contaminants and additives, pollutants and pesticides.
For detection of medium to large biomolecules, such as nucleic acids, proteins, membranes and their components, both intrinsic and extrinsic schemes have been used. The advantage of an intrinsic method is that the Raman signature will reflect the state of the analyte of interest, but researchers must keep in mind that the binding to the metal surface can in itself denature, or at least change, a protein to some extent. To achieve SERS from these molecules, it is sometimes possible to functionalize the surface of the SERS substrate that enables the capture of a specific analyte molecule. The advantage of an extrinsic method, in which the target biomolecule is labeled or a SERS tag is introduced, is access to much lower detection limits.
Specific nucleic acid sequences can be detected by hybridization with a complementary target. The ability to detect specific nucleic acids can be important in molecular diagnostics, such as the identification of viruses and bacteria and detection of disease states. The process also aids in the identification of criminal suspects. Target antigens or antibodies can be detected using immunoassay platforms and have been applied to drug identification, food quality control, environmental monitoring and clinical diagnostics. In one scheme, in order to achieve lower limits of detection, a sandwich assay technique has been used in which single-walled carbon nanotubes, which provide enormous Raman signals, were conjugated to antibodies designed to capture a specific analyte and detect down to femtomolar concentrations. In fact, the simultaneous use of different antibodies enabled the simultaneous detection of different target proteins.
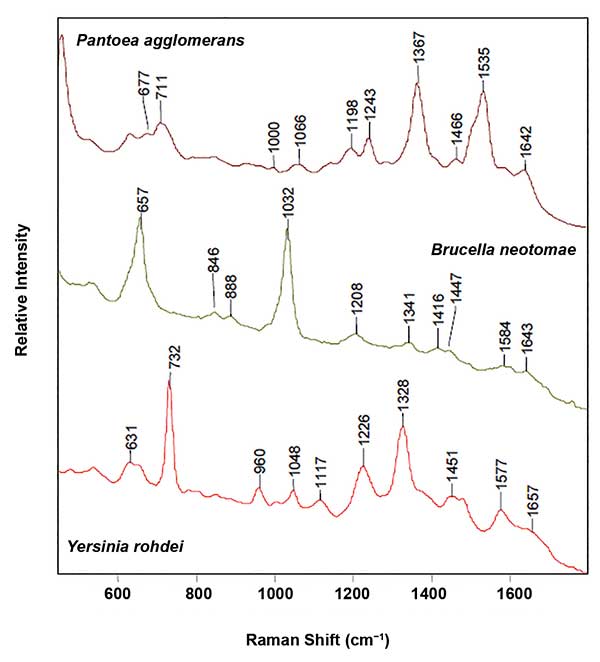 SERS spectra of three gram-negative bacteria: P. agglomerans, B. neotomae and Y. rohdei, illustrating the sensitivity of SERS to speciation. Reproduced with permission from J. Guicheteau and S.D. Christesen (2006). Principal component analysis of bacteria using surface-enhanced Raman spectroscopy, Proc SPIE, Vol. 6218.
SERS spectra of three gram-negative bacteria: P. agglomerans, B. neotomae and Y. rohdei, illustrating the sensitivity of SERS to speciation. Reproduced with permission from J. Guicheteau and S.D. Christesen (2006). Principal component analysis of bacteria using surface-enhanced Raman spectroscopy, Proc SPIE, Vol. 6218.
One of the most important applications of SERS is the detection and identification of bacteria. This capability has the potential to assist in emergent care where identification of the most effective antibiotic for an infected individual usually takes two to three days. For a patient in sepsis, this delay can be fatal. Unlike the SERS signal of many molecules, the SERS signals of bacteria are not the same as the normal Raman signals of the same materials. That is because of the SERS sensitivity to the exterior of the bacteria that bind to the SERS substrate. To utilize the SERS signal for bacterial identification and differentiation between strains, multivariate analysis techniques are an absolute necessity, as is an extensive database. Implicit in this is the reproducibility of the SERS substrates or particles.
SERS and cancer
SERS is being explored for cancer diagnosis, especially in its early stages14. Its application will depend on the identification of multiple biomarkers for various malignant diseases. In addition, as the malignancy progresses, its markers probably change. As an example of the potential power of this technology, researchers have used SERS to identify patients with esophageal cancer by analyzing blood serum15.
SERS has also been utilized to assist with theranostics, which is the integration of diagnostics and therapeutics. Delivery of a photoactive drug to a malignant cell produces a highly toxic reactive oxygen species when exposed to light. However, because the concentrations have to be controlled in order to prevent damage to healthy tissue, SERS can be useful for monitoring those drug amounts. Additionally, the metal nanoparticles themselves can be used for photodynamic therapy.
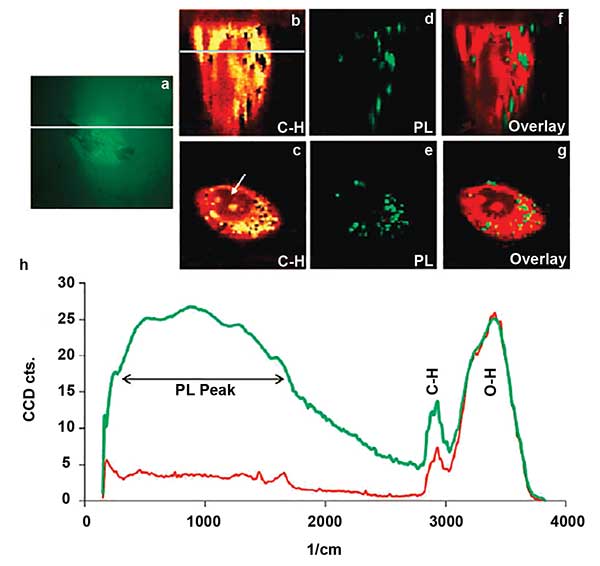
Confocal Raman microscope image of single cell. An optical image of cell, with the line indicating where depth scanning was performed (a). A depth scan of the cell using the intensity of the CH band (b). The line indicates the depth at which the lateral scan (c) was performed. Depth and lateral images constructed from the photoluminescence peak generated at the gold particles (d) and (e). An overlay of the depth (b and d) and lateral (c and e) images (f) and (g). The chart (h) is representative of Raman spectra of the cell (red) and gold nanoparticles (green). Reprinted with permission from Neha B. Shah, Jinping Dong and John C. Bischof (2011). Cellular uptake and nanoscale localization of gold nanoparticles in cancer using label-free confocal Raman microscopy. Molecular Pharmaceutics, Vol. 81, pp. 176-184.
One of the holy grails of SERS is its use to define tumor margins where histology does not identify all of the transformed tissue at the edges of the growth. Application of SERS in the operating room is not yet a reality, but tests are ongoing in animal models. This application shows promise for improving surgical outcomes for diseases such as glioblastoma, which has a recurrence rate of 80 to 90 percent. The use of a Raman endoscopic probe is essential for developing use of SERS diagnostics in the operating room theater16. Normal Raman has been implemented for gastrointestinal endoscopy but measurement of SERS after administration of nanoparticles is being pursued17.
Meet the authors
Fran Adar is a Raman principal scientist with Horiba Instruments Inc. in Edison, N.J.; email: [email protected]. Maruda
Shanmugasundaram is an AFM-Raman applications scientist with Horiba; email: [email protected].
References
1. For a simple introduction to plasmonics of SERS substrates, see: F. Adar (2015). SERS: An update of progress made. Spectroscopy, Vol. 30, pp. 14-19.
2. M. Moskovits (1985). Surfaced-enhanced spectroscopy. Rev Modern Phys, Vol. 57, Issue 3, pp. 783-826.
3. A.M. Michaels et al. (1999). Surface enhanced Raman spectroscopy of individual rhodamine 6G molecules on large Ag nanocrystals, J Am Chem Soc, Vol. 121, pp. 9932-9939.
4. N.J. Halas et al. (2011). Plasmons in strongly coupled metallic nanostructures. Chemical Reviews, Vol. 111, pp. 3913-3961.
5. M. Procházka (2016). Surface-Enhanced Raman Spectrocopy: Bioanalytical, Biomolecular and Medical Applications. Cham, Switzerland: Springer International Publishing.
6. Ibid, 30.
7. S. Schlücker, ed. (2011). Surface Enhanced Raman Spectroscopy: Analytical, Biophysical and Life Science Applications. Weinheim, Germany: Wiley-VCH, Chapter 8.
8. Halas.
9. M.J. Natan (2006). Concluding remarks. Surface enhanced Raman scattering, Faraday Discuss, Vol. 132, pp. 321-328.
10. Procházka, p. 68.
11. Schlücker, Chapter 12.
12. S. Dochow et al. (2013). Raman-on-chip device and detection fibres with fibre Bragg grating for analysis of solutions and particles. Lab Chip, Vol. 13, pp. 1109-1114.
13. Schlücker, Chapter 8.
14. Procházka, Chapter 7.
15. X. Li et al. (2015). Detecting esophageal cancer using surface-enhanced Raman spectroscopy (SERS) of serum coupled with hierarchical cluster analysis and principal component analysis, Appl Spectros, Vol. 69, Issue 11, pp. 1334-1341.
16. Dochow.
17. Procházka, p. 198.
Tip: enhanced Raman spectroscopy (TERS), which will be discussed this summer in another BioPhotonics article by Adar and Shanmugasundaram, involves the use of a fine metal tip to produce surface-enhancing conditions with a spatial resolution that is consistent with scanning probe microscopes (SPMs). TERS can act as a bridge between diffraction-limited, far-field optical techniques, such as SERS and high-resolution topographic imaging techniques of the SPMs, by correlating chemical and morphological features at the nanoscale level. The forthcoming article will review how TERS can be used to elucidate outstanding biological and medical questions.