Accurate and predictive software modeling could allow standardization for small-animal imaging.
Silas Leavesley, Dr. Bartek Rajwa and Dr. J. Paul Robinson, Purdue University,and Dr. Edward Freniere, Richard Hassler and Linda Smith, Lambda Research Corp.
Small animals represent vital model systems for the study of organism development and human disease. They are increasingly recognized as powerful discovery tools in genetic, proteomic and cancer research. For example, animal studies have vast utility in studying efficacy and toxicity of novel cancer therapeutics.
Until recently, the significant potential of animal studies was untapped because it was necessary to sacrifice the animals to perform tissue or molecular analysis. Once an animal is sacrificed, it is not possible for researchers or clinicians to observe the natural or perturbed evolution of the processes under investigation. Now, functional, molecular and morphologic quantitative imaging techniques provide data about the biochemical, genetic and pharmacological processes in the same animal. These in vivo studies can be performed noninvasively and in real time. However, a lack of standards and calibration references significantly reduces the value of these imaging techniques.
Small-animal imaging depends on the ability of visible and near-infrared wavelengths to noninvasively penetrate small distances through living tissues. There are many versions of commercially available small-animal imagers, and all face a common set of complications associated with imaging through a medium that is simultaneously absorptive, scattering and fluorescent.
Imaging a weak fluorescence signal within this environment can be complicated. Trying to infer any type of radiometric or quantitative measurement from these images is very difficult. Clinical medical imaging modalities such as x-ray, CT, PET and ultrasound face many of the same problems as fluorescence small-animal imagers. Imaging through tissues is overcome by the penetrating nature of the types of radiation and target tissues. The quality and control factor, however, is not yet overcome.
Clinical imaging systems employ a standard schedule of quality control and validation that allows them to yield consistent and, often, calibrated imaging data. Phantom reference standards are the tools most commonly used in performing this quality control and validation. Unlike other clinical imaging techniques, a reproducible and informative phantom does not exist today in small-animal fluorescence imaging.
Establishing a standard among competing in vivo fluorescence imagers would bring the benefits of interoperability. The ability to analyze data interchangeably from multiple imaging systems and from multiple clinical trials and experiments facilitates adoption and would allow exponentially more and thus less expensive data to be available for analysis.
Fluorescent phantoms
One of the most significant problems with designing fluorescent phantoms is that the combined scattering, absorption and fluorescence phenomena make straightforward mathematical calculation of phantom performance difficult for flat-plane geometries and nearly impossible for anything more complex. For example, it is possible to combine multiple types of phantom materials, such as machining plastic phantom material that can contain various fluorescent solutions or materials, but developing a mathematical model that predicts the response of this phantom is extremely difficult. Because of this, development of a fluorescence tissue phantom has been achieved mostly by empirical techniques.
Researchers at Purdue University’s Bindley Bioscience Center have developed models of absorptive, scattering and fluorescent phantoms that are used to predict the response in small-animal fluorescence imagers. The models were developed with TracePro software, a tool from Lambda Research Corp. that models and analyzes the propagation of light in imaging and illumination optomechanical systems.
Modeling the distribution of fluorophore concentration across tissue types and modeling environmental factors such as the physiological animal positioning require flexible solid modeling capability, the ability to characterize fluorescent probes, and the ability to import measured laboratory data into the software model. Furthermore, to simultaneously account for multiple system parameters — sensitivity, quantitative accuracy, multiplex capability and tissue depth — the software must simulate dependencies of all optical phenomena without approximation to achieve accurate predictive models.
The model used for design and analysis of the phantom took into account a fluorescence imager, mouse samples and the phantom. The software performed system-level modeling and analysis of light distributions, stray light, throughput, flux absorbed by surfaces and bulk media, and fluorescence and polarization effects.
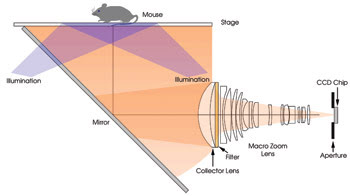
Figure 1. To design a fluorescent phantom for use with small-animal imaging, all imaging parts had to be simulated. This is the optical configuration of a fluorescence imager that was simulated.
The fluorescence imager model was created by importing mechanics and imaging optics from commercially available mechanical and lens design software (Figure 1). The luminous flux and numerical aperture of the fiber optic delivery systems for the multiple excitation sources (515, 568, 621, 674, 715, 745, 775 and 1580 nm) were modeled, as were the spectral response and active area of the CCD. Finally, fluorophore-specific fluorescence filter sets were added to the model.
To model the mouse, Micro CT datasets from individual mice were imported into GE Healthcare’s MicroView software (Figure 2), which defined surface meshes delimiting the edges of skin, bones and tissue. The data was exported as stereo lithography files to 3-D StudioMax from Autodesk Inc., where the anatomical structures were smoothed, and meshes were made “watertight.” The edited meshes were exported in a file exchange format and imported into TracePro, where they were checked for geometric and mesh errors and then stitched together to form a solid object.
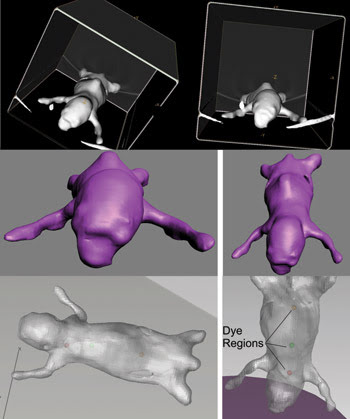
Figure 2. To simulate the mouse, the animal’s CT isosurface was rendered in MicroView (top); then the CT mesh was smoothed in 3D Studio Max (middle), and finally the solid model was displayed in TracePro with fluorescent dye regions (bottom).
With the object solids defined, the bulk absorption, scatter and fluorescence properties were specified from TracePro material databases to mimic skin, bone and tissue. Three 1-mm-diameter spheres were created to simulate fluorophores injected into the mouse. The fluorescent pockets were modeled by specifying the fluorescence properties, including the concentration and quantum efficiency of the injected dyes.
Optical properties of the fluorophores — including relative excitation and emission curves and peak molar extinction coefficients — were retrieved from the Invitrogen Molecular Probes Products catalog resident in TracePro. Optical properties of the sample tissues were retrieved from the software’s biological tissue catalog.
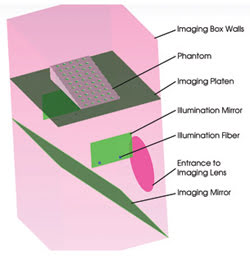
Figure 3. The phantom model is represented in TracePro.
The phantom must have scattering and absorption properties that are characteristic of small-animal (mouse) tissues. Polyester, polyurethane and silicone phantoms are permanent matrix compositions that are suitable for routine calibration and testing. Common scattering materials are lipid-based emulsions, titanium or aluminum oxide powders, and polymer microspheres. The fluorescence absorbers are the same molecular dyes used in the small animal for tagging. All these materials, including the commercially available dyes, were imported from the TracePro database or from measured material data.
One fluorescence phantom design was based on absorption phantoms used in x-ray imaging that employ a series of progressively thicker x-ray-absorbing plates to test the penetration depth of the x-ray. For optical imaging, a series of progressively thicker optically absorbing and scattering material was used. A high concentration of each fluorescent dye (Alexa Fluor 532, 610, 660, 680, 750) was added to the phantom via a series of 50-μl wells in the phantom (Figure 4). A different dye was added to each column, so that there was one dye of each species at each thickness of absorptive and scattering material. The solid geometry was created in CAD software and imported into TracePro, where optical material properties for each object were specified.

Figure 4. A fluorescent phantom with successively thicker layers of absorbing and scattering material and wells containing five fluorescent dyes was used.
Simulation and analysis
To simulate the whole system, the mouse and phantom models can be placed into the modeled environment of a small-animal fluorescence imager. Ray traces then may be executed where excitation source rays propagate through the complete system model with portions of the flux of each ray allocated to absorption, specular reflection and transmission, fluorescence, polarization and scattering (Figure 5).

Figure 5. Ray traces can be executed where excitation source rays propagate through the complete system model.
The simulations allow comparison of mouse and phantom models in regard to system throughput at the CCD, flux absorbed by object surfaces and bulk material, and stray and scattered light from mechanical, optical and biological materials.
The primary advantage of using this advanced modeling software in phantom development is that design alternatives for the phantom may be modeled and tested without the cost and time associated with iterative hardware prototyping and laboratory and clinical testing. For example, by characterizing and adding the fluorescent properties of the plastic materials used to construct the phantom, their contribution to autofluorescence may be isolated and accounted for in the calibration reference.
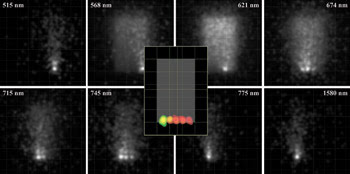
Figure 6. The irradiance maps of fluorescence emission are shown as a function of wavelength (outer gray-scale images) for the fluorescence phantom model. At each wavelength, a different fluorophore has its peak emission. Ray-trace sampling, and scattering and autofluorescence from the phantom, contribute to perceived blur. A false-color overlay of irradiance maps at individual fluorescence peak emission wavelengths (central insert) shows the location of dyes with respect to the outlined phantom.
Autofluorescence interferes with detection of the resulting specific fluorescent signals and is caused by fluorescence from structures other than the dye-tagged target. The concentration and quantum efficiency of the dye and/or the materials and geometry of the phantom also may be modified to achieve simulations that are repeatable and that support more accurate simulations performed with actual small-animal models.
The modeling of fluorescence excitation, emission and propagation within living tissue has been a limiting factor in small-animal diagnostic imaging. The ability to simulate animals and tissue phantoms in model imagers with sufficient statistical samples and calibration references will allow the development of standards and thus foster widespread deployment of small-animal imaging.
Meet the authors
Silas Leavesley is a doctoral student at the Bindley Bioscience Center of Purdue University in West Lafayette, Ind.
Bartek Rajwa is a postdoctoral research associate at Purdue University.
J. Paul Robinson is a professor at Purdue University and deputy director for cytomics and imaging at the Bindley Bioscience Center.
Edward Freniere is president of Lambda Research Corp. in Littleton, Mass.
Richard Hassler is executive vice president of Lambda Research Corp.
Linda Smith is vice president of sales and marketing at Lambda Research Corp.; e-mail: [email protected].