
3-D model reveals cancer cell motility
Raquel Harper
Most cancers become fatal when cells
break away from the primary tumor and travel through the bloodstream to other parts
of the body, where they settle and divide once again. Pharmaceutical companies use
2-D assays to investigate drugs that can combat cancer cell migration. In these
assays, researchers have found that the cells move across the surface of a matrix
in a single plane using counteracting tractile and adhesion forces. However, 3-D
confocal images now have shown that something different might be happening.
Muhammad H. Zaman and his colleagues, working
at Whitehead Institute for Biomedical Research in Cambridge, Mass., believed that
investigations on artificial 2-D plates might not provide all the crucial information
about cell motility. According to Zaman, understanding how cancer cells migrate
through the bloodstream will enable researchers to determine which individual receptors
are responsible for the cancer cells’ increased migration. He believes that
they can target those receptors and find a way to control metastasis.
Zaman and his team decided to look
at the trends of prostate cancer cells’ movement in the 3-D environment of
gels derived from mouse sarcomas. Because cancer cells use clawlike proteins called
integrins to pull themselves forward through their surrounding matrix, Zaman disabled
some of these integrins to see what the 3-D imaging would reveal.
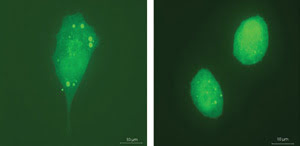
Researchers have discovered that 3-D imaging reveals information
about a cancer cell’s motility that 2-D imaging does not. The confocal image
on the left shows a prostate cancer cell in 3-D with all of its integrins intact
and expressing. The right 3-D image shows the change in a cancer cell’s shape
when some of its integrins are blocked, which means that the cell’s ability
to migrate also is different. Images reprinted with permission of PNAS.
They labeled the cells with a green
tracker dye from Molecular Probes (now Invitrogen Corp.) and embedded them inside
the gel to mimic a tissue environment. After the gel polymerized, they imaged the
cells with a confocal microscope — from PerkinElmer Inc. of Wellesley, Mass.
— with a spinning disk at 25x and 40x magnifications.
Zaman explained that only a confocal
instrument could provide high-resolution imaging that would allow them to look at
the details of one cell at a time as well as of a population of cancer cells over
a long period. Bright-field microscopy would only enable them to image the whole
field at once, resulting in photobleaching after just a few hours.
The researchers collected images every
15 minutes for six hours, with each image consisting of a 100-μm Z-stack at
0.5-μm intervals. They also collected 2-D data on all of the same cancer cells,
so that they could provide a good comparison with their 3-D results. Zaman said
that this was perhaps the most challenging part of the experiment, as it took them
a long time to acquire both 2- and 3-D data.
He had been working simultaneously
on a mathematical model that would help explain how drugs influence metastasis.
The model predicted that the density and stiffness of the extracellular matrix would
significantly affect a cancer cell’s motility.
The imaging revealed contrasting results
to the well-established findings from 2-D assays. Instead of compensating for a
decrease in the number of integrins by binding in areas of higher matrix densities
(as found in 2-D systems), the cells in 3-D appeared to find areas that were less
dense. They also took on a different shape as they moved through pores of varying
sizes.
Zaman believes that their findings
help explain why 2-D assays for metastasis-inhibiting drugs do not effectively predict
their effects in tissue. He hopes to see 3-D assays, accompanied by appropriate
computational models, used to predict how drugs will affect metastasis.
Now at the University of Texas in Austin,
Zaman plans to continue investigating how the cell attaches to the matrix and, specifically,
to find which receptors are responsible for cancer cell migration. He also would
like to study cancer cells that come directly from the patient, instead of those
that have been being transformed to cell culture.
PNAS, July 18, 2006, pp. 10889-10894.
Published: September 2006